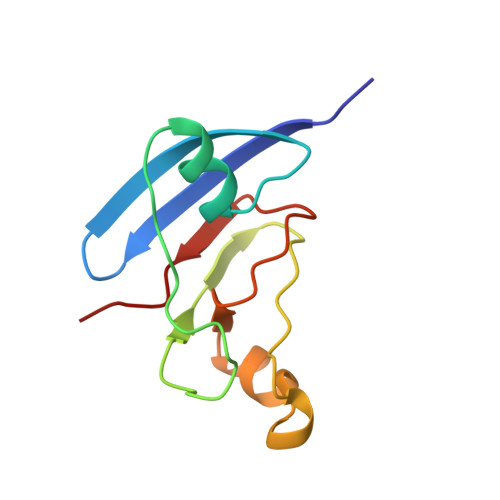
NMR structure of the [2Fe-2S] ferredoxin domain from soluble methane monooxygenase reductase and interaction with its hydroxylase.
Muller, J., Lugovskoy, A.A., Wagner, G., Lippard, S.J.(2002) Biochemistry 41: 42-51
- PubMed: 11772001 Search on PubMed
- DOI: https://doi.org/10.1021/bi015668k
- Primary Citation Related Structures:
1JQ4 - PubMed Abstract:
The soluble methane monooxygenase (sMMO) from Methylococcus capsulatus (Bath) is a multicomponent enzyme system required for the conversion of methane to methanol. It comprises a hydroxylase, a regulatory protein, and a reductase. The reductase contains two domains: an NADH-binding and FAD-containing flavin domain and a ferredoxin (Fd) domain carrying a [2Fe-2S] cofactor. Here, we report the solution structure of the reduced form of the 98-amino acid Fd domain (Blazyk, J. L., and Lippard, S. J. Unpublished results) determined by nuclear magnetic resonance (NMR) spectroscopy and restrained molecular dynamics calculations. The structure consists of six beta strands arranged into two beta sheets as well as three alpha helices. Two of these helices form a helix-proline-helix motif, unprecedented among [2Fe-2S] proteins. The [2Fe-2S] cluster is coordinated by the sulfur atoms of cysteine residues 42, 47, 50, and 82. The 10.9 kDa ferredoxin domain of the reductase protein transfers electrons to carboxylate-bridged diiron centers in the 251 kDa hydroxylase component of sMMO. The binding of the Fd domain with the hydroxylase was investigated by NMR spectroscopy. The hydroxylase binding surface on the ferredoxin protein has a polar center surrounded by patches of hydrophobic residues. This arrangement of amino acids differs from that by which previously studied [2Fe-2S] proteins interact with their electron-transfer partners. The critical residues on the Fd domain involved in this binding interaction map well onto the universally conserved residues of sMMO enzymes from different species. We propose that the [2Fe-2S] domains in these other sMMO systems have a fold very similar to the one found here for M. capsulatus (Bath) MMOR-Fd.
- Department of Chemistry, Massachusetts Institute of Technology, Cambridge, Massachusetts 02139-4307, USA.
Organizational Affiliation: