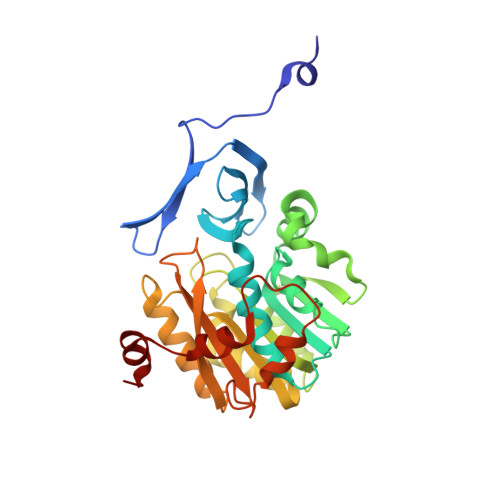
The crystal structure of spermidine synthase with a multisubstrate adduct inhibitor.
Korolev, S., Ikeguchi, Y., Skarina, T., Beasley, S., Arrowsmith, C., Edwards, A., Joachimiak, A., Pegg, A.E., Savchenko, A.(2002) Nat Struct Biol 9: 27-31
- PubMed: 11731804 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nsb737
- Primary Citation Related Structures:
1INL, 1JQ3 - PubMed Abstract:
Polyamines are essential in all branches of life. Spermidine synthase (putrescine aminopropyltransferase, PAPT) catalyzes the biosynthesis of spermidine, a ubiquitous polyamine. The crystal structure of the PAPT from Thermotoga maritima (TmPAPT) has been solved to 1.5 A resolution in the presence and absence of AdoDATO (S-adenosyl-1,8-diamino-3-thiooctane), a compound containing both substrate and product moieties. This, the first structure of an aminopropyltransferase, reveals deep cavities for binding substrate and cofactor, and a loop that envelops the active site. The AdoDATO binding site is lined with residues conserved in PAPT enzymes from bacteria to humans, suggesting a universal catalytic mechanism. Other conserved residues act sterically to provide a structural basis for polyamine specificity. The enzyme is tetrameric; each monomer consists of a C-terminal domain with a Rossmann-like fold and an N-terminal beta-stranded domain. The tetramer is assembled using a novel barrel-type oligomerization motif.
- Biosciences Division and Structural Biology Center, Argonne National Laboratory, 9700 South Cass Ave., Bldg. 202, Argonne, Illinois 60439, USA.
Organizational Affiliation: