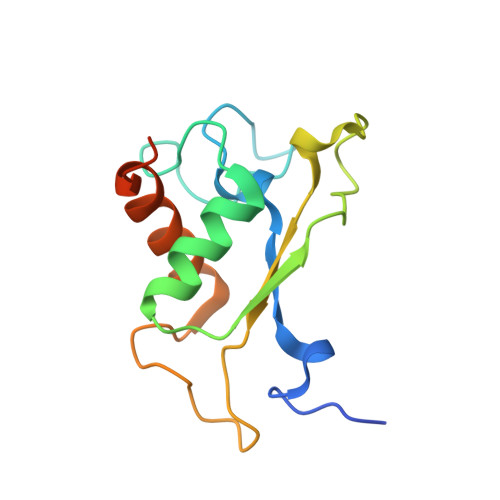
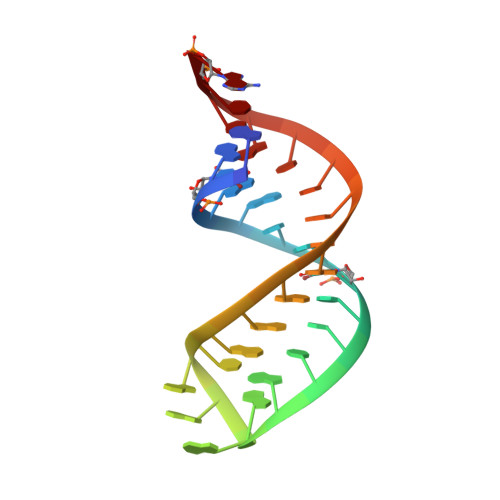
Crystal structure of an early protein-RNA assembly complex of the signal recognition particle.
Wild, K., Sinning, I., Cusack, S.(2001) Science 294: 598-601
- PubMed: 11641499 Search on PubMed
- DOI: https://doi.org/10.1126/science.1063839
- Primary Citation Related Structures:
1JID - PubMed Abstract:
The signal recognition particle (SRP) is a universally conserved ribonucleoprotein complex that mediates the cotranslational targeting of secretory and membrane proteins to cellular membranes. A crucial early step in SRP assembly in archaea and eukarya is the binding of protein SRP19 to specific sites on SRP RNA. Here we report the 1.8 angstrom resolution crystal structure of human SRP19 in complex with its primary binding site on helix 6 of SRP RNA, which consists of a stem-loop structure closed by an unusual GGAG tetraloop. Protein-RNA interactions are mediated by the specific recognition of a widened major groove and the tetraloop without any direct protein-base contacts and include a complex network of highly ordered water molecules. A model of the assembly of the SRP core comprising SRP19, SRP54, and SRP RNA based on crystallographic and biochemical data is proposed.
- Biochemie-Zentrum (BZH), University of Heidelberg, Im Neuenheimer Feld 328, D-69120 Heidelberg, Germany. klemens.wild@bzh.uni-heidelberg.de
Organizational Affiliation: