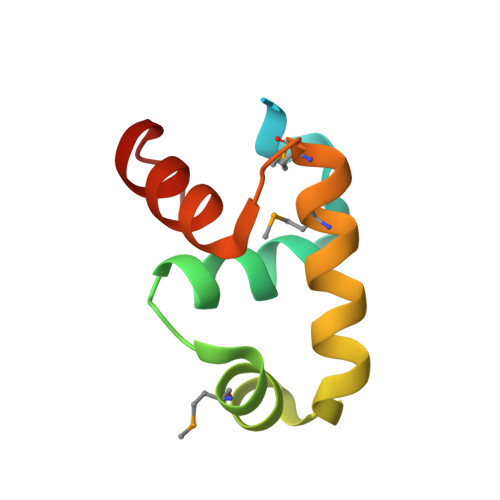
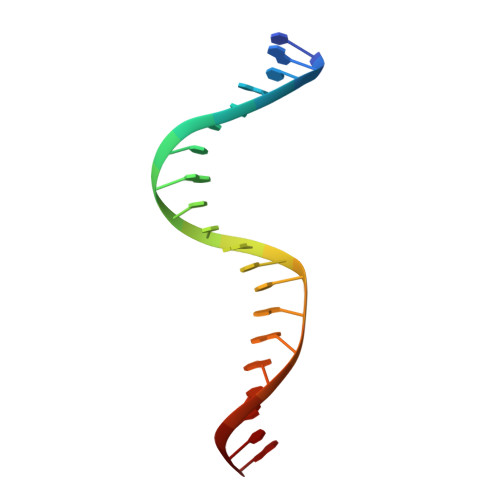
Dimerization allows DNA target site recognition by the NarL response regulator.
Maris, A.E., Sawaya, M.R., Kaczor-Grzeskowiak, M., Jarvis, M.R., Bearson, S.M., Kopka, M.L., Schroder, I., Gunsalus, R.P., Dickerson, R.E.(2002) Nat Struct Biol 9: 771-778
- PubMed: 12352954 Search on PubMed
- DOI: https://doi.org/10.1038/nsb845
- Primary Citation Related Structures:
1JE8 - PubMed Abstract:
Two-component signal transduction systems are modular phosphorelay regulatory pathways common in prokaryotes. In the co-crystal structure of the Escherichia coli NarL signal output domain bound to DNA, we observe how the NarL family of two-component response regulators can bind DNA. DNA recognition is accompanied by the formation of a new dimerization interface, which could occur only in the full-length protein via a large intramolecular domain rearrangement. The DNA is recognized by the concerted effects of solvation, van der Waals forces and inherent DNA deformability, rather than determined primarily by major groove hydrogen bonding. These subtle forces permit a small DNA-binding domain to perturb the DNA helix, leading to major DNA curvature and a transition from B- to A-form DNA at the binding site, where valine on the recognition helix interacts unexpectedly with the polar major groove floor.
- Department of Chemistry and Biochemistry, University of California, Los Angeles, P.O. Box 951569, Los Angeles, California 90095-1569, USA.
Organizational Affiliation: