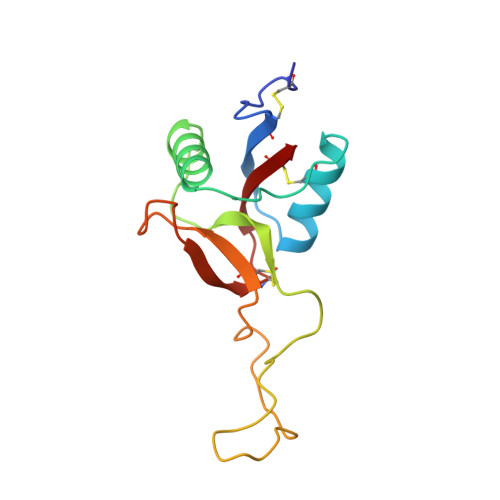
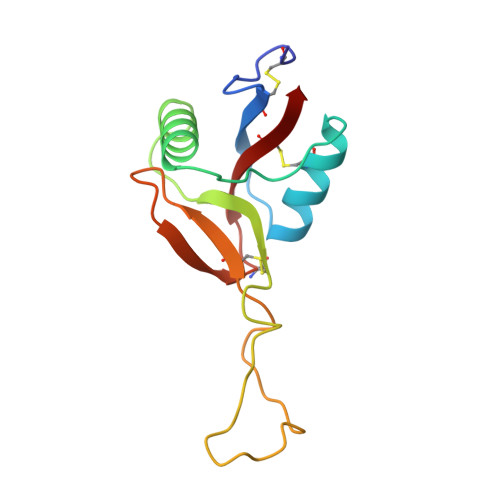
Structure of coagulation factors IX/X-binding protein, a heterodimer of C-type lectin domains.
Mizuno, H., Fujimoto, Z., Koizumi, M., Kano, H., Atoda, H., Morita, T.(1997) Nat Struct Biol 4: 438-441
- PubMed: 9187649 Search on PubMed
- DOI: https://doi.org/10.1038/nsb0697-438
- Primary Citation Related Structures:
1IXX - PubMed Abstract:
Coagulation factors IX/X-binding protein is an intertwined dimer with a central loop projecting into the adjoining subunit. Excluding this loop, each subunit has a fold similar to rat mannose-binding protein.