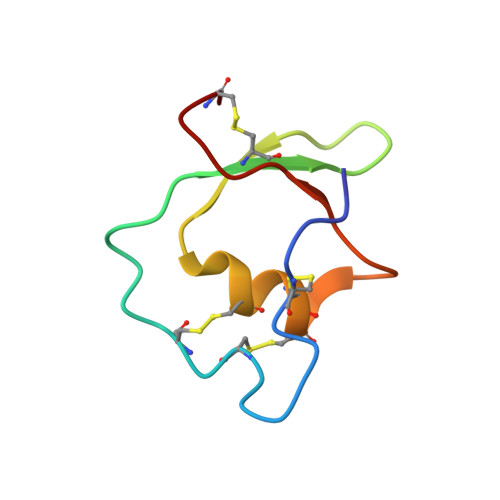
Solution structure of ascidian trypsin inhibitor determined by nuclear magnetic resonance spectroscopy.
Hemmi, H., Yoshida, T., Kumazaki, T., Nemoto, N., Hasegawa, J., Nishioka, F., Kyogoku, Y., Yokosawa, H., Kobayashi, Y.(2002) Biochemistry 41: 10657-10664
- PubMed: 12186551 Search on PubMed
- DOI: https://doi.org/10.1021/bi026035o
- Primary Citation Related Structures:
1IW4 - PubMed Abstract:
The three-dimensional solution structure of ascidian trypsin inhibitor (ATI), a 55 amino acid residue protein with four disulfide bridges, was determined by means of two-dimensional nuclear magnetic resonance (2D NMR) spectroscopy. The resulting structure of ATI was characterized by an alpha-helical conformation in residues 35-42 and a three-stranded antiparallel beta-sheet in residues 22-26, 29-32, and 48-50. The presence of an alpha-helical conformation was predicted from the consensus sequences of the cystine-stabilized alpha-helical (CSH) motif, which is characterized by an alpha-helix structure in the Cys-X(1)-X(2)-X(3)-Cys portion (corresponding to residues 37-41), linking to the Cys-X-Cys portion (corresponding to residues 12-14) folded in an extended structure. The secondary structure and the overall folding of the main chain of ATI were very similar to those of the Kazal-type inhibitors, such as Japanese quail ovomucoid third domain (OMJPQ3) and leech-derived tryptase inhibitor form C (LDTI-C), although ATI does not show extensive sequence homology to these inhibitors except for a few amino acid residues and six of eight half-cystines. On the basis of these findings, we realign the amino acid sequences of representative Kazal-type inhibitors including ATI and discuss the unique structure of ATI with four disulfide bridges.
- National Food Research Institute, 2-1-12 Kannondai, Tsukuba, Ibaraki 305-8642, Japan.
Organizational Affiliation: