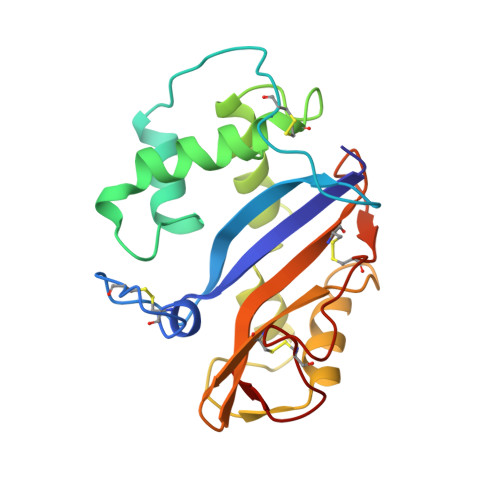
Crystal structure at 1.5-A resolution of Pyrus pyrifolia pistil ribonuclease responsible for gametophytic self-incompatibility.
Matsuura, T., Sakai, H., Unno, M., Ida, K., Sato, M., Sakiyama, F., Norioka, S.(2001) J Biological Chem 276: 45261-45269
- PubMed: 11577107 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M107617200
- Primary Citation Related Structures:
1IQQ - PubMed Abstract:
The crystal structure of the Pyrus pyrifolia pistil ribonuclease (S(3)-RNase) responsible for gametophytic self-incompatibility was determined at 1.5-A resolution. It consists of eight helices and seven beta-strands, and its folding topology is typical of RNase T(2) family enzymes. Based on a structural comparison of S(3)-RNase with RNase Rh, a fungal RNase T(2) family enzyme, the active site residues of S(3)-RNase assigned were His(33) and His(88) as catalysts and Glu(84) and Lys(87) as stabilizers of an intermediate in the transition state. Moreover, amino acid residues that constitute substrate binding sites of the two RNases could be superimposed geometrically. A hypervariable (HV) region that has an S-allele-specific sequence comprises a long loop and short alpha-helix. This region is far from the active site cleft, exposed on the molecule's surface, and positively charged. Four positively selected (PS) regions, in which the number of nonsynonymous substitutions exceeds that of synonymous ones, are located on either side of the active site cleft, and accessible to solvent. These structural features suggest that the HV or PS regions may interact with a pollen S-gene product(s) to recognize self and non-self pollen.
- Institute for Protein Research, Osaka University, Suita, Osaka 565-0871, Japan.
Organizational Affiliation: