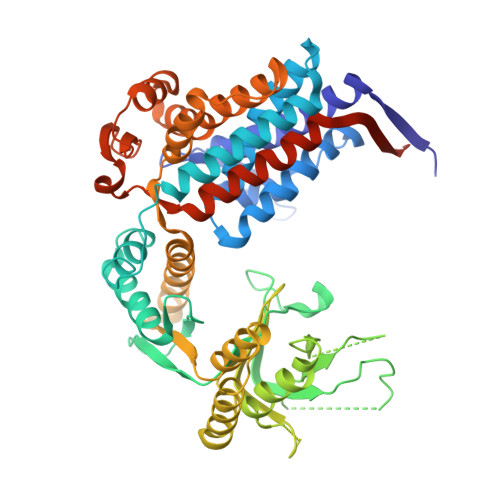
Crystal structure of chaperonin-60 from Paracoccus denitrificans.
Fukami, T.A., Yohda, M., Taguchi, H., Yoshida, M., Miki, K.(2001) J Mol Biology 312: 501-509
- PubMed: 11563912 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.2001.4961
- Primary Citation Related Structures:
1IOK - PubMed Abstract:
The crystal structure of chaperonin-60 from Paracoccus denitrificans (P.cpn60) has been determined at 3.2 A resolution by the molecular replacement method. Two heptameric rings of identical subunits of P.cpn60 in adjacent asymmetric units are stacked in a back-to-back manner and form a cylinder, as found in GroEL, cpn60 from Escherichia coli. With respect to the unliganded GroEL structure, each subunit of P.cpn60 tilts 2 degrees outwards and the apical domain twists 4 degrees counter-clockwise in the top view in a hinge-like manner, rendering the central hole 5 A wider. Despite the subunit tilts, both rings in P.cpn60 contact at two sites of the equatorial domain in the same way as in GroEL. Interactions between residues 434 and 434, and 463 and 463 observed in GroEL were not found in P.cpn60, and the interaction between 452 and 461 was weaker in P.cpn60 than in GroEL. The unique hydrogen bond between 468 and 471 was observed at the right site in P.cpn60, which could account for why the subunits tilt outwards. The contact surface area was reduced at the left site, which is similar to the observed changes in the GroEL structures induced by ATP binding. In general, inter-ring interactions in P.cpn60 were weakened, which is consistent with findings that P.cpn60 is observed in single-ring forms as well as in double-ring forms.
- Department of Chemistry Graduate School of Science, Kyoto University, Kyoto, Sakyo-ku, 606-8502, Japan.
Organizational Affiliation: