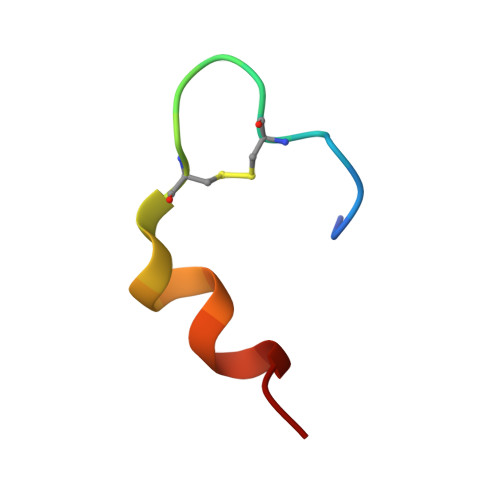
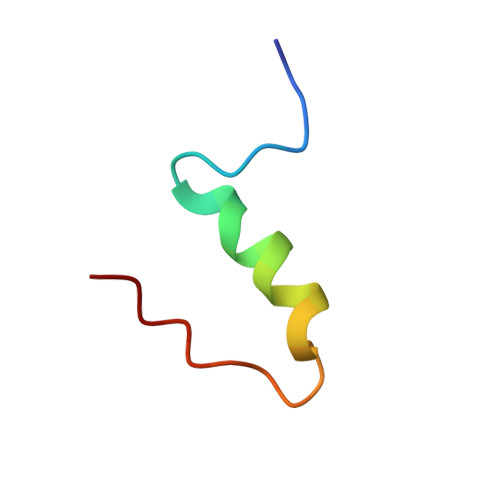
The relationship between insulin bioactivity and structure in the NH2-terminal A-chain helix.
Olsen, H.B., Ludvigsen, S., Kaarsholm, N.C.(1998) J Mol Biology 284: 477-488
- PubMed: 9813131 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1998.2175
- Primary Citation Related Structures:
1IOG, 1IOH - PubMed Abstract:
Studies of naturally occuring and chemically modified insulins have established that the NH2-terminal helix of the A-chain is important in conferring affinity in insulin-receptor interactions. Nevertheless, the three-dimensional structural basis for these observations has not previously been studied in detail. To correlate structure and function in this region of the molecule, we have used the solution structure of an engineered monomer (GluB1, GluB10, GluB16, GluB27, desB30)-insulin (4E insulin) as a template for design of A-chain mutants associated with enhanced or greatly diminished affinity for the insulin receptor. In the context of 4E insulin, the employed mutants, i.e. ThrA8-->His and ValA3-->Gly, result in species with 143% and 0.1% biological activity, respectively, relative to human insulin. The high-resolution NMR studies reveal two well-defined structures each resembling the template. However, significant structural differences are evident notably in residues A2-A8 and their immediate environment. In comparison with the template structure, the A8His mutation enhances the helical character of residues A2-A8. This structural change leads to additional exposure of a hydrophobic patch mainly consisting of species invariant residues. In contrast, the A3Gly mutation leads to stretching and disruption of the A2-A8 helix and changes both the dimensions and the access to the hydrophobic patch exposed in the more active insulins. We conclude that the mutations induce small, yet decisive structural changes that either mediate or inhibit the subtle conformational adjustments involved in the presentation of this part of the insulin pharmacophore to the receptor.
- Novo Nordisk A/S, Novo Alle 1, Bagsvaerd, DK-2880, Denmark.
Organizational Affiliation: