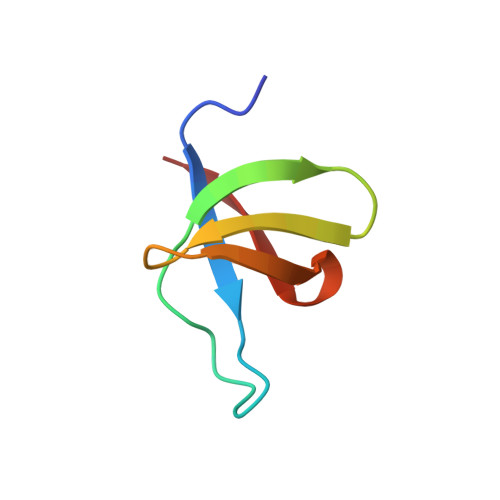
Solution structure of the DNA binding domain of HIV-1 integrase.
Lodi, P.J., Ernst, J.A., Kuszewski, J., Hickman, A.B., Engelman, A., Craigie, R., Clore, G.M., Gronenborn, A.M.(1995) Biochemistry 34: 9826-9833
- PubMed: 7632683 Search on PubMed
- DOI: https://doi.org/10.1021/bi00031a002
- Primary Citation Related Structures:
1IHV, 1IHW - PubMed Abstract:
The solution structure of the DNA binding domain of HIV-1 integrase (residues 220-270) has been determined by multidimensional NMR spectroscopy. The protein is a dimer in solution, and each subunit is composed of a five-stranded beta-barrel with a topology very similar to that of the SH3 domain. The dimer is formed by a stacked beta-interface comprising strands 2, 3, and 4, with the two triple-stranded antiparallel beta-sheets, one from each subunit, oriented antiparallel to each other. One surface of the dimer, bounded by the loop between strands beta 1 and beta 2, forms a saddle-shaped groove with dimensions of approximately 24 x 23 x 12 A in cross section. Lys264, which has been shown from mutational data to be involved in DNA binding, protrudes from this surface, implicating the saddle-shaped groove as the potential DNA binding site.
- Laboratory of Chemical Physics, National Institute of Diabetes and Digestive and Kidney Diseases, National Institutes of Health, Bethesda, Maryland 20892-0520, USA.
Organizational Affiliation: