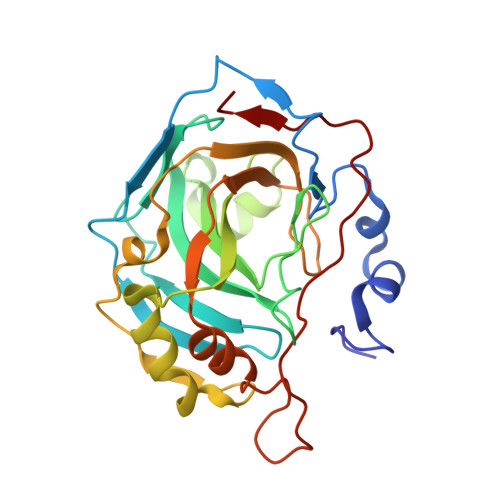
Combinatorial computational method gives new picomolar ligands for a known enzyme.
Grzybowski, B.A., Ishchenko, A.V., Kim, C.Y., Topalov, G., Chapman, R., Christianson, D.W., Whitesides, G.M., Shakhnovich, E.I.(2002) Proc Natl Acad Sci U S A 99: 1270-1273
- PubMed: 11818565 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.032673399
- Primary Citation Related Structures:
1IF7, 1IF8, 1IF9 - PubMed Abstract:
Combinatorial small molecule growth algorithm was used to design inhibitors for human carbonic anhydrase II. Two enantiomeric candidate molecules were predicted to bind with high potency (with R isomer binding stronger than S), but in two distinct conformations. The experiments verified that computational predictions concerning the binding affinities and the binding modes were correct for both isomers. The designed R isomer is the best-known inhibitor (K(d) approximately 30 pM) of human carbonic anhydrase II.
- Harvard University, Department of Chemistry and Chemical Biology, 12 Oxford Street, Cambridge, MA 02138, USA.
Organizational Affiliation: