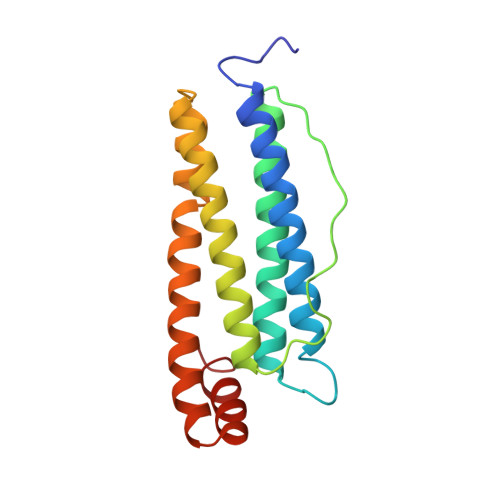
Comparison of the structures of the cubic and tetragonal forms of horse-spleen apoferritin.
Granier, T., Gallois, B., Dautant, A., Langlois d'Estaintot, B., Precigoux, G.(1997) Acta Crystallogr D Biol Crystallogr 53: 580-587
- PubMed: 15299889 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444997003314
- Primary Citation Related Structures:
1IER, 1IES - PubMed Abstract:
Horse-spleen apoferritin is known to crystallize in three different space groups, cubic F432, tetragonal P42(1)2 and orthorhombic P2(1)2(1)2. A structure comparison of the cubic and tetragonal forms is presented here. Both crystal forms were obtained by the vapor-diffusion technique and data were collected at 2.26 A (cubic crystal) and 2.60 A (tetragonal crystal) resolution. Two main differences were observed between these crystal structures: (i) whereas intermolecular contacts only involve salt-bridge type interactions via cadmium ions in the cubic structure, two types of interactions are observed in the tetragonal crystal (cadmium-ion-mediated salt bridges and hydrogen-bonding interactions) and (ii) cadmium ions bound in the threefold axes of ferritin molecules exhibit lower site-occupation factors in the tetragonal structure than in the cubic one.
- Unité de Biophysique Structurale, EP534 CNRS, Université Bordeaux 1, Talence, France.
Organizational Affiliation: