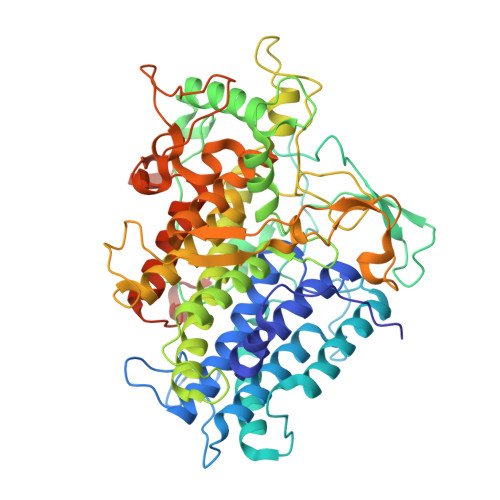
Implications for the catalytic mechanism of the vanadium-containing enzyme chloroperoxidase from the fungus Curvularia inaequalis by X-ray structures of the native and peroxide form.
Messerschmidt, A., Prade, L., Wever, R.(1997) Biol Chem 378: 309-315
- PubMed: 9165086 Search on PubMed
- DOI: https://doi.org/10.1515/bchm.1997.378.3-4.309
- Primary Citation Related Structures:
1IDQ, 1IDU - PubMed Abstract:
Implications for the catalytic mechanism of the vanadium-containing chloroperoxidase from the fungus Curvularia inaequalis have been obtained from the crystal structures of the native and peroxide forms of the enzyme. The X-ray structures have been solved by difference Fourier techniques using the atomic model of the azide chloroperoxidase complex. The 2.03 A crystal structure (R = 19.7%) of the native enzyme reveals the geometry of the intact catalytic vanadium center. The vanadium is coordinated by four non-protein oxygen atoms and one nitrogen (NE2) atom from histidine 496 in a trigonal bipyramidal fashion. Three oxygens are in the equatorial plane and the fourth oxygen and the nitrogen are at the apexes of the bipyramid. In the 2.24 A crystal structure (R = 17.7%) of the peroxide derivate the peroxide is bound to the vanadium in an eta2-fashion after the release of the apical oxygen ligand. The vanadium is coordinated also by 4 non-protein oxygen atoms and one nitrogen (NE2) from histidine 496. The coordination geometry around the vanadium is that of a distorted tetragonal pyramid with the two peroxide oxygens, one oxygen and the nitrogen in the basal plane and one oxygen in the apical position. A mechanism for the catalytic cycle has been proposed based on these X-ray structures and kinetic data.
- Max-Planck-Institut für Biochemie, Martinsried bei München, Germany.
Organizational Affiliation: