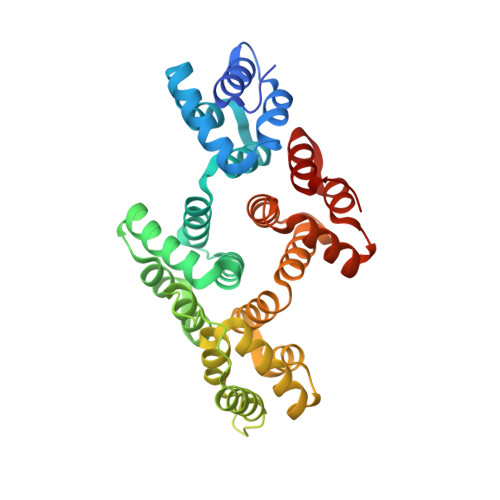
Phosphorylation mutants elucidate the mechanism of annexin IV-mediated membrane aggregation.
Kaetzel, M.A., Mo, Y.D., Mealy, T.R., Campos, B., Bergsma-Schutter, W., Brisson, A., Dedman, J.R., Seaton, B.A.(2001) Biochemistry 40: 4192-4199
- PubMed: 11300800 Search on PubMed
- DOI: https://doi.org/10.1021/bi002507s
- Primary Citation Related Structures:
1I4A - PubMed Abstract:
Site-directed mutagenesis, electron microscopy, and X-ray crystallography were used to probe the structural basis of annexin IV-induced membrane aggregation and the inhibition of this property by protein kinase C phosphorylation. Site-directed mutants that either mimic (Thr6Asp, T6D) or prevent (Thr6Ala, T6A) phosphorylation of threonine 6 were produced for these studies and compared with wild-type annexin IV. In vitro assays showed that unmodified wild-type annexin IV and the T6A mutant, but not PKC-phosphorylated wild-type or the T6D mutant, promote vesicle aggregation. Electron crystallographic data of wild-type and T6D annexin IV revealed that, similar to annexin V, the annexin IV proteins form 2D trimer-based ordered arrays on phospholipid monolayers. Cryo-electron microscopic images of junctions formed between lipid vesicles in the presence of wild-type annexin IV indicated a separation distance corresponding to the thickness of two layers of membrane-bound annexin IV. In this orientation, a single layer of WT annexin IV, attached to the outer leaflet of one vesicle, would undergo face-to-face self-association with the annexin layer of a second vesicle. The 2.0-A resolution crystal structure of the T6D mutant showed that the mutation causes release of the N-terminal tail from the protein core. This change would preclude the face-to-face annexin self-association required to aggregate vesicles. The data suggest that reversible complex formation through phosphorylation and dephosphorylation could occur in vivo and play a role in the regulation of vesicle trafficking following changes in physiological states.
- Departments of Molecular and Cellular Physiology and of Obstetrics and Gynecology, University of Cincinnati, College of Medicine, Ohio 45220, USA.
Organizational Affiliation: