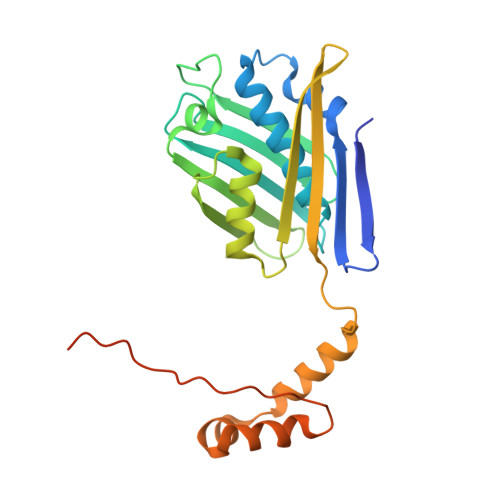
The 2.2 A crystal structure of Hsp33: a heat shock protein with redox-regulated chaperone activity.
Vijayalakshmi, J., Mukhergee, M.K., Graumann, J., Jakob, U., Saper, M.A.(2001) Structure 9: 367-375
- PubMed: 11377197 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(01)00597-4
- Primary Citation Related Structures:
1HW7 - PubMed Abstract:
One strategy that cells employ to respond to environmental stresses (temperature, oxidation, and pathogens) is to increase the expression of heat shock proteins necessary to maintain viability. Several heat shock proteins function as molecular chaperones by binding unfolded polypeptides and preventing their irreversible aggregation. Hsp33, a highly conserved bacterial heat shock protein, is a redox-regulated molecular chaperone that appears to protect cells against the lethal effects of oxidative stress. The 2.2 A crystal structure of a truncated E. coli Hsp33 (residues 1-255) reveals a domain-swapped dimer. The core domain of each monomer (1-178) folds with a central helix that is sandwiched between two beta sheets. The carboxyl-terminal region (179-235), which lacks the intact Zn binding domain of Hsp33, folds into three helices that pack on the other subunit. The interface between the two core domains is comprised of conserved residues, including a rare Glu-Glu hydrogen bond across the dyad axis. Two potential polypeptide binding sites that span the dimer are observed: a long groove containing pockets of conserved and hydrophobic residues, and an intersubunit 10-stranded beta sheet "saddle" with a largely uncharged or hydrophobic surface. Hsp33 is a dimer in the crystal structure. Solution studies confirmed that this dimer reflects the structural changes that occur upon activation of Hsp33 as a molecular chaperone. Patterns of conserved residues and surface charges suggest that two grooves might be potential binding sites for protein folding intermediates.
- Biophysics Research Division, University of Michigan, 48109, Ann Arbor, MI, USA.
Organizational Affiliation: