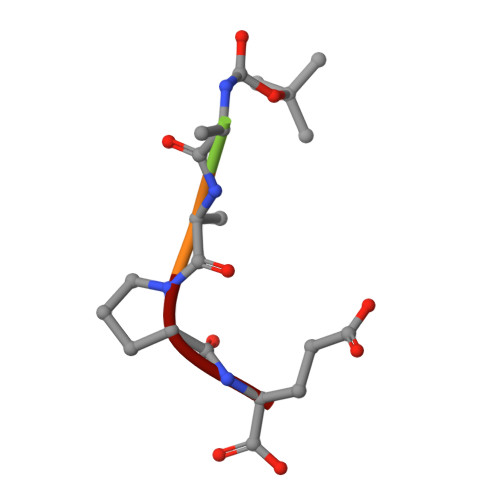
A glutamic acid specific serine protease utilizes a novel histidine triad in substrate binding.
Nienaber, V.L., Breddam, K., Birktoft, J.J.(1993) Biochemistry 32: 11469-11475
- PubMed: 8105890 Search on PubMed
- DOI: https://doi.org/10.1021/bi00094a001
- Primary Citation Related Structures:
1HPG - PubMed Abstract:
Proteases specific for cleavage after acidic residues have been implicated in several disease states, including epidermolysis, inflammation, and viral processing. A serine protease with specificity toward glutamic acid substrates (Glu-SGP) has been crystallized in the presence of a tetrapeptide ligand and its structure determined and refined to an R-factor of 17% at 2.0-A resolution. This structure provides an initial description of the design of proteolytic specificity for negatively charged residues. While the overall fold of Glu-SGP closely resembles that observed in the pancreatic-type serine proteases, stabilization of the negatively charged substrate when bound to this protein appears to involve a more extensive part of the protease than previously observed. The substrate carboxylate is bound to a histidine side chain, His213, which provides the primary electrostatic compensation of the negative charge on the substrate, and to two serine hydroxyls, Ser192 and Ser216. Glu-SGP displays maximum activity at pH 8.3, and assuming normal pKa's, the glutamate side chain and His213 will be negatively charged and neutral, respectively, at this pH. In order for His213 to carry a positive charge at the optimal pH, its pKa will have to be raised by at least two units. An alternative mechanism for substrate charge compensation is suggested that involves a novel histidine triad, His213, His199, and His228, not observed in any other serine protease. The C-terminal alpha-helix, ubiquitous to all pancreatic-type proteases, is directly linked to this histidine triad and may also play a role in substrate stabilization.(ABSTRACT TRUNCATED AT 250 WORDS)
- Department of Biochemistry and Molecular Biophysics, Washington University School of Medicine, St. Louis, Missouri 63110.
Organizational Affiliation: