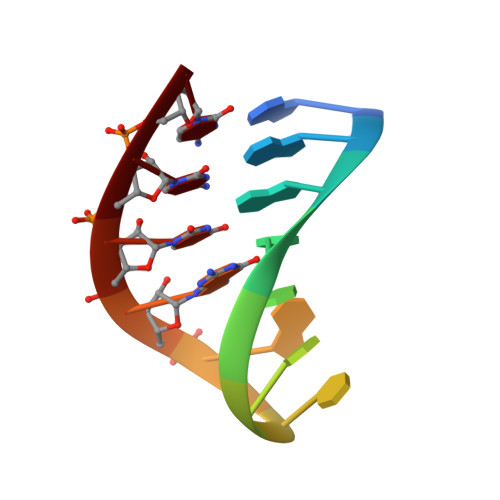
Solution structure of an arabinonucleic acid (ANA)/RNA duplex in a chimeric hairpin: comparison with 2'-fluoro-ANA/RNA and DNA/RNA hybrids.
Denisov, A.Y., Noronha, A.M., Wilds, C.J., Trempe, J.F., Pon, R.T., Gehring, K., Damha, M.J.(2001) Nucleic Acids Res 29: 4284-4293
- PubMed: 11691916 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/29.21.4284
- Primary Citation Related Structures:
1HO6, 1HOQ - PubMed Abstract:
Hybrids of RNA and arabinonucleic acid (ANA) as well as the 2'-fluoro-ANA analog (2'F-ANA) were recently shown to be substrates of the enzyme RNase H. Although RNase H binds to double-stranded RNA, no cleavage occurs with such duplexes. Therefore, knowledge of the structure of ANA/RNA hybrids may prove helpful in the design of future antisense oligonucleotide analogs. In this study, we have determined the NMR solution structures of ANA/RNA and DNA/RNA hairpin duplexes and compared them to the recently published structure of a 2'F-ANA/RNA hairpin duplex. We demonstrate here that the sugars of RNA nucleotides of the ANA/RNA hairpin stem adopt the C3'-endo (north, A-form) conformation, whereas those of the ANA strand adopt a 'rigid' O4'-endo (east) sugar pucker. The DNA strand of the DNA/RNA hairpin stem is flexible, but the average DNA/RNA hairpin structural parameters are close to the ANA/RNA and 2'F-ANA/RNA hairpin parameters. The minor groove width of ANA/RNA, 2'F-ANA/RNA and DNA/RNA helices is 9.0 +/- 0.5 A, a value that is intermediate between that of A- and B-form duplexes. These results rationalize the ability of ANA/RNA and 2'F-ANA/RNA hybrids to elicit RNase H activity.
- Department of Biochemistry and Montreal Joint Centre for Structural Biology, McGill University, Montreal, QC H3G 1Y6, Canada.
Organizational Affiliation: