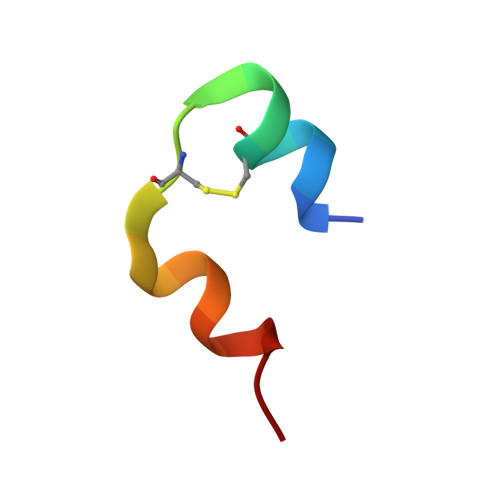
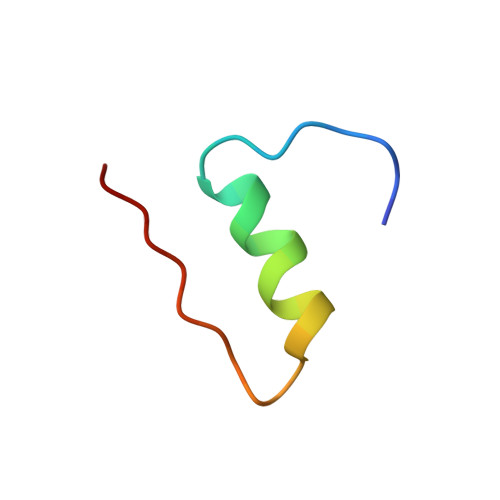
High-resolution structure of an engineered biologically potent insulin monomer, B16 Tyr-->His, as determined by nuclear magnetic resonance spectroscopy.
Ludvigsen, S., Roy, M., Thogersen, H., Kaarsholm, N.C.(1994) Biochemistry 33: 7998-8006
- PubMed: 8025104 Search on PubMed
- DOI: https://doi.org/10.1021/bi00192a003
- Primary Citation Related Structures:
1HLS - PubMed Abstract:
Site-directed mutagenesis is used in conjunction with 1H nuclear magnetic resonance (NMR) and circular dichroism (CD) spectroscopy in order to find an insulin species amenable for structure determination in aqueous solution by NMR spectroscopy. A successful candidate in this respect, i.e., B16 Tyr-->His mutant insulin, is identified and selected for detailed characterization by two-dimensional 1H NMR. This mutant species retains 43% biological potency and native folding stability, but in contrast to human insulin it remains monomeric at millimolar concentration in aqueous solution at pH 2.4. The resulting homogeneous sample allows high-quality 2D NMR spectra to be recorded. The NMR studies result in an almost complete assignment of the 1H resonance signals as well as identification of NOE cross peaks. NOE-derived distance restraints in conjunction with torsion restraints based on measured coupling constants, 3JHNH alpha, are used for structure calculations using the hybrid method of distance geometry and simulated annealing. The calculated structures show that the major part of the insulin monomer is structurally well-defined with an average rms deviation between the 20 calculated structures and the mean coordinates of 0.89 A for all backbone atoms, 0.46 A for backbone atoms (A2-A19 and B4-B28), and 1.30 A for all heavy atoms. The structure of the A-chain is composed of two helices from A2 to A7 and from A12 to A19 connected by a short extended strand. The B-chain consists of a loop, B1-B8, an alpha-helix, B9-B19, a beta-turn, B20-B23, and an extended strand from B24 to B30.(ABSTRACT TRUNCATED AT 250 WORDS)
- Novo Research Institute, Novo Nordisk A/S, Bagsvaerd, Denmark.
Organizational Affiliation: