The Structure of the Coliphage Hk022 Nun Protein-Lambda-Phage Boxb RNA Complex. Implications for the Mechanism of Transcription Termination
Faber, C., Schaerpf, M., Becker, T., Sticht, H., Roesch, P.(2001) J Biological Chem 276: 32064
- PubMed: 11356847 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M102975200
- Primary Citation Related Structures:
1HJI - PubMed Abstract:
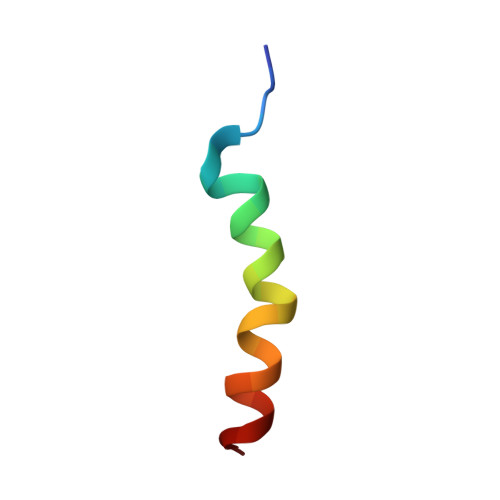
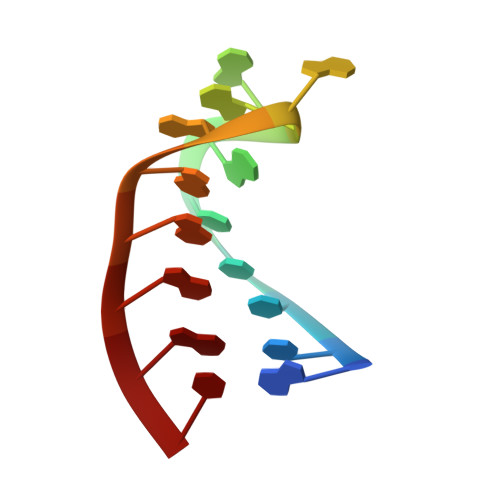
Nun protein from coliphage HK022 binds to phage boxB RNA and functions, in contrast to phage lambda N protein, as a transcriptional terminator. The basic Nun-(10-44) peptide contains the boxB RNA binding arginine rich motif, ARM. The peptide binds boxB RNA and competes with the phage lambda ARM peptide N-(1-36) as indicated by nuclear magnetic resonance (NMR) spectroscopy titrations. In two-dimensional nuclear Overhauser enhancement spectroscopy experiments boxB RNA in complex with Nun-(20-44) exhibits the same pattern of resonances as it does in complex with N peptides containing the ARM, and we could show that Nun-(20-44) forms a bent alpha-helix upon binding to the boxB RNA. The structure of the boxB RNA-bound Nun-(20-44) was determined on the basis of 191 intra- and 30 intermolecular distance restraints. Ser-24 is anchored to the lower RNA stem, and stacking of Tyr-39 and A7 is clearly experimentally indicated. Arg-28 shows numerous contacts to the RNA stem. Leu-22, Ile-30, Trp-33, Ile-37, and Leu-41 form a hydrophobic surface, which could be a recognition site for additional host factors such as NusG. Such a hydrophobic surface area is not present in N-(1-36) bound to boxB RNA.
- Lehrstuhl für Biopolymere, Universität Bayreuth, Universitätsstr. 30, 95440 Bayreuth, Germany.
Organizational Affiliation: