The High Resolution Crystal Structure of Alpha-Conotoxin Si to 0.75 Angstroms
Janes, R.W., Hargittai, B., Barany, G., Maclean, E.J., Teat, S.J.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| ALPHA-CONOTOXIN SI | 14 | Conus striatus | Mutation(s): 0 | 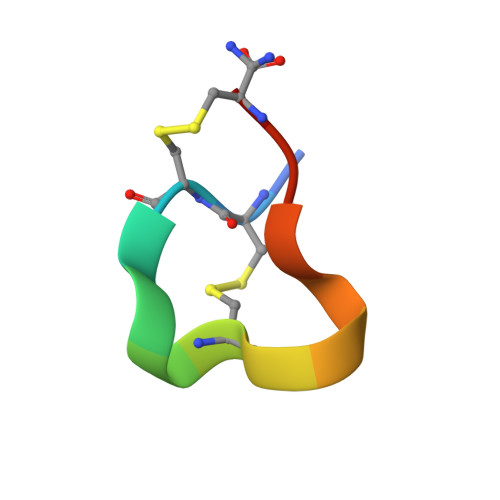 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P15471 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 32.092 | α = 90 |
| b = 32.092 | β = 90 |
| c = 14.669 | γ = 120 |
| Software Name | Purpose |
|---|---|
| SHELXL-97 | refinement |
| SAINT | data reduction |
| SADABS | data scaling |
| SHELX | phasing |