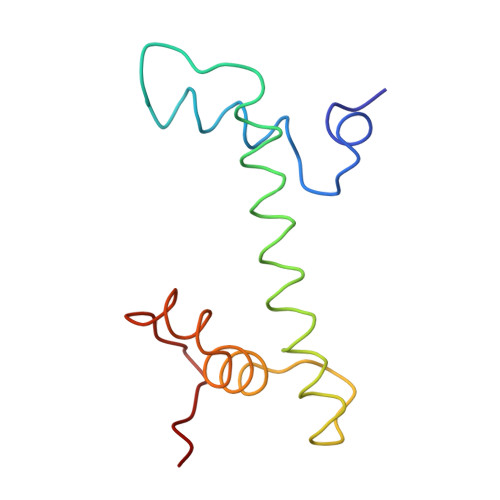
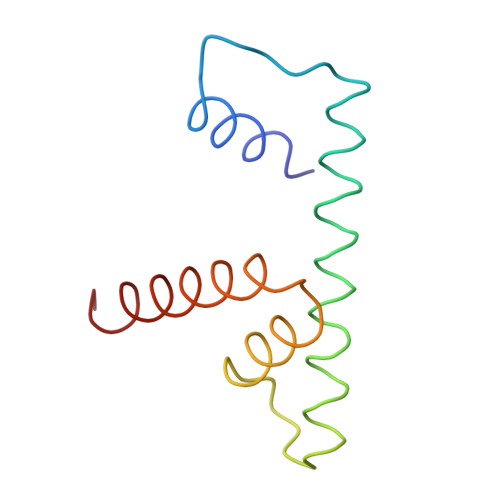
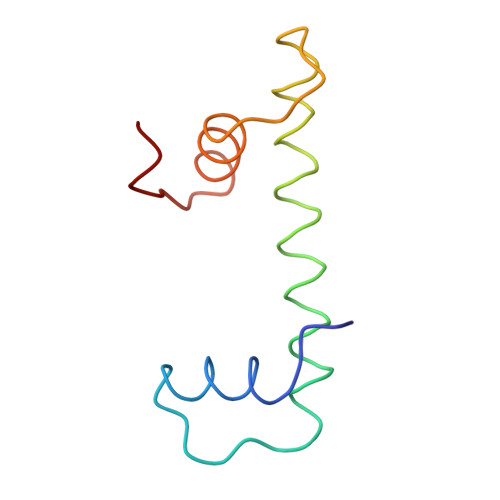
The nucleosomal core histone octamer at 3.1 A resolution: a tripartite protein assembly and a left-handed superhelix.
Arents, G., Burlingame, R.W., Wang, B.C., Love, W.E., Moudrianakis, E.N.(1991) Proc Natl Acad Sci U S A 88: 10148-10152
- PubMed: 1946434 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.88.22.10148
- Primary Citation Related Structures:
1HIO, 2HIO - PubMed Abstract:
The structure of the octameric histone core of the nucleosome has been determined by x-ray crystallography to a resolution of 3.1 A. The histone octamer is a tripartite assembly in which a centrally located (H3-H4)2 tetramer is flanked by two H2A-H2B dimers. It has a complex outer surface; depending on the perspective, the structure appears as a wedge or as a flat disk. The disk represents the planar projection of a left-handed proteinaceous superhelix with approximately 28 A pitch. The diameter of the particle is 65 A and the length is 60 A at its maximum and approximately 10 A at its minimum extension; these dimensions are in agreement with those reported earlier by Klug et al. [Klug, A., Rhodes, D., Smith, J., Finch, J. T. & Thomas, J. O. (1980) Nature (London) 287, 509-516]. The folded histone chains are elongated rather than globular and are assembled in a characteristic "handshake" motif. The individual polypeptides share a common central structural element of the helix-loop-helix type, which we name the histone fold.
- Department of Biology, Johns Hopkins University, Baltimore, MD 21218.
Organizational Affiliation: