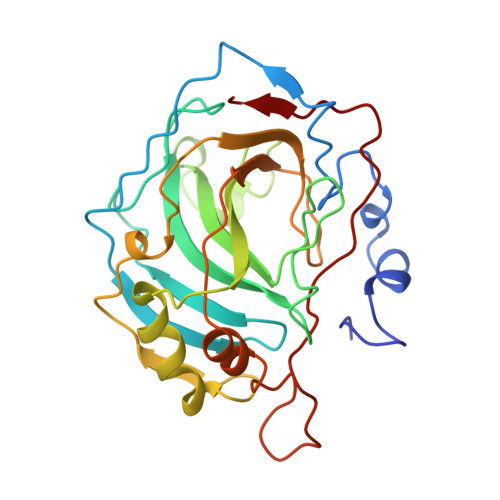
Structural consequences of hydrophilic amino acid substitutions in the hydrophobic pocket of human carbonic anhydrase II.
Nair, S.K., Christianson, D.W.(1993) Biochemistry 32: 4506-4514
- PubMed: 8485129 Search on PubMed
- DOI: https://doi.org/10.1021/bi00068a005
- Primary Citation Related Structures:
1HEA, 1HEB, 1HEC, 1HED - PubMed Abstract:
The three-dimensional structures of Leu-198-->Glu, Leu-198-->His, Leu-198-->Arg, and Leu-198-->Ala variants of human carbonic anhydrase II (CAII) have each been determined by X-ray crystallographic methods to a resolution of 2.0 A. The side chain of Leu-198 is located at the mouth of the active site hydrophobic pocket, and this pocket is required for substrate association. Hydrophobic-->hydrophilic amino acid substitutions at the mouth of the pocket decrease kcat/KM for CO2 hydration: the CO2 hydrase activities of Leu-198-->Glu, Leu-198-->His, and Leu-198-->Arg CAIIs are diminished 19-fold, 10-fold, and 17-fold, respectively, relative to the wild-type enzyme; however, the substitution of a compact aliphatic side chain for Leu-198 has a smaller effect on catalysis, in that Leu-198-->Ala CAII exhibits only a 3-fold decrease in CO2 hydrase activity [Krebs, J. F., Rana, F., Dluhy, R. A., & Fierke, C. A. (1993) Biochemistry (preceding paper in this issue)]. It is intriguing that CO2 hydrase activity is not severely diminished in Leu-198-->Arg CAII, even though the side chain of Arg-198 blocks the hydrophobic pocket. Therefore, the bulky side chain of Arg-198 must be reasonably mobile in order to accommodate substrate association. Significantly, a residue larger than the wild-type Leu-198 side chain does not necessarily block the substrate association pocket; e.g., the side chain of Glu-198 packs against a hydrophobic patch, the net result of which is a wider mouth for the pocket.(ABSTRACT TRUNCATED AT 250 WORDS)
- Department of Chemistry, University of Pennsylvania, Philadelphia 19104-6323.
Organizational Affiliation: