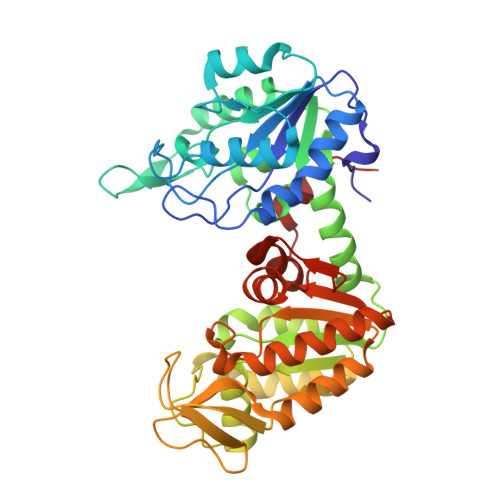
A 1.8 A resolution structure of pig muscle 3-phosphoglycerate kinase with bound MgADP and 3-phosphoglycerate in open conformation: new insight into the role of the nucleotide in domain closure.
Szilagyi, A.N., Ghosh, M., Garman, E., Vas, M.(2001) J Mol Biology 306: 499-511
- PubMed: 11178909 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.2000.4294
- Primary Citation Related Structures:
1HDI - PubMed Abstract:
3-phosphoglycerate kinase (PGK) is a typical kinase with two structural domains. The domains each bind one of the two substrates, 3-phosphoglycerate (3-PG) and MgATP. For the phospho-transfer reaction to take place the substrates must be brought closer by a hinge-bending domain closure. Open and closed structures of the enzyme with different relative domain positions have been determined from different species, but a comprehensive description of this conformational transition is yet to be attained. Crystals of pig muscle PGK in complex with MgADP and 3-phosphoglycerate were grown under the conditions which have previously resulted in crystals of the closed, catalytically competent conformation of Trypanosoma brucei PGK. The X-ray structure of the pig muscle ternary complex was determined at 1.8 A and the model was refined to R=20.8% and Rfree=24.1%. Contrary to expectation, however, it represents an essentially open conformation compared to that of T. brucei PGK. In addition, the beta-phosphate group of ADP is mobile in the new structure, in contrast to its well-defined position in T. brucei PGK. An extensive comparison of the ternary complexes from these remote species has been carried out in order to establish general differences between the two conformations and is reported here. A second pair of the open and closed structures was also compared. These analyses have made it possible to define several characteristic changes which accompany the structural transition, in addition to those identified previously: (1) the operation of a hinge at beta-strand L in the inter-domain region which greatly affects the relative domain positions; (2) the rearrangement and movement of helix 8, regulated through the interactions with the nucleotide phosphate; and (3) the existence of another hinge between helix 14 and the rest of the C-terminal part of the chain, which allows fine adjustment of the N-domain position. The main hinge at beta-strand L acts in concert with the C-terminal hinge at helix 7 described previously. Simultaneous interactions of the nucleotide phosphate groups with the loop that precedes helix 8, beta-strand J and the N terminus of helix 13 are required for propagation of the nucleotide effect towards the beta-strand L molecular hinge. A detailed description of the role of nucleotide binding in the hinge operation is presented.
- Institute of Enzymology Biological Research Center, Hungarian Academy of Sciences, Budapest, H-1518, Hungary.
Organizational Affiliation: