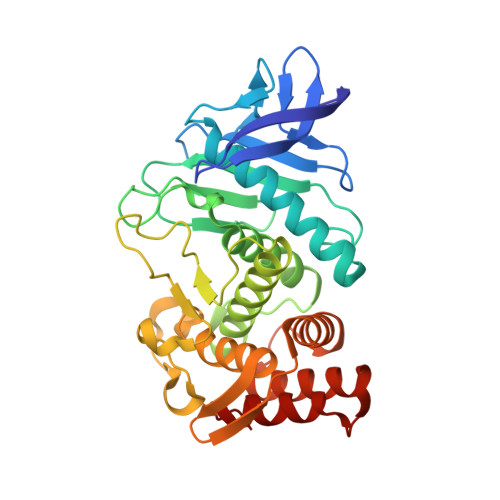
The 2.2 A Resolution Structure of Thermolysin (Tln) Crystallized in the Presence of Potassium Thiocyanate.
Gaucher, J., Selkti, M., Prange, T., Tomas, A.(2002) Acta Crystallogr D Biol Crystallogr 58: 2198
- PubMed: 12454500 Search on PubMed
- DOI: https://doi.org/10.1107/s0907444902015457
- Primary Citation Related Structures:
1GXW - PubMed Abstract:
A new crystallization protocol for thermolysin (EC 3.4.24.27) from Bacillus thermoproteolyticus is presented. After dissolving the protein in the presence of KSCN, which avoids the use of DMSO and CsCl, crystals were obtained following the salting-in method. Crystal cell parameters are isomorphous with those previously reported from DMSO/CsCl mixtures. The new SCN(-) crystal structure has been analyzed. It shows the presence of one thiocyanate ion in the catalytic site and several rearrangements in the S(1) and S(2) subsites. These results are in agreement with the measurements of Inouye et al. [(1998), J. Biochem. (Tokyo), 123, 847-852], who observed in solution that the solubility of TLN, which is particularly poor in low ionic strength solutions, increases dramatically in the presence of several neutral salts. The results reported here suggest possible explanations for the solubility increase and for the inhibitory effects of high SCN(-) concentrations on thermolysin activity.
- Laboratoire de Cristallographie et RMN Biologiques (UMR-8015, CNRS), Université Paris V, Faculté de Pharmacie, 4 Avenue de l'Observatoire, 75270 Paris CEDEX 06, France. gaucher@pharmacie.univ-paris5.fr
Organizational Affiliation: