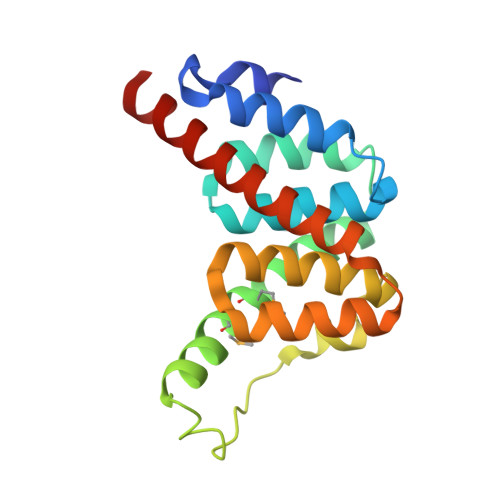
The Crystal Structure of Mycobacterium Tuberculosis Alkylhydroperoxidase Ahpd, a Potential Target for Antitubercular Drug Design
Nunn, C.M., Djordjevic, S., Hillas, P.J., Nishida, C., Ortiz de Montellano, P.R.(2002) J Biological Chem 277: 20033
- PubMed: 11914371 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M200864200
- Primary Citation Related Structures:
1GU9 - PubMed Abstract:
The resistance of Mycobacterium tuberculosis to isoniazid is commonly linked to inactivation of a catalase-peroxidase, KatG, that converts isoniazid to its biologically active form. Loss of KatG is associated with elevated expression of the alkylhydroperoxidases AhpC and AhpD. AhpD has no sequence identity with AhpC or other proteins but has alkylhydroperoxidase activity and possibly additional physiological activities. The alkylhydroperoxidase activity, in the absence of KatG, provides an important antioxidant defense. We have determined the M. tuberculosis AhpD structure to a resolution of 1.9 A. The protein is a trimer in a symmetrical cloverleaf arrangement. Each subunit exhibits a new all-helical protein fold in which the two catalytic sulfhydryl groups, Cys-130 and Cys-133, are located near a central cavity in the trimer. The structure supports a mechanism for the alkylhydroperoxidase activity in which Cys-133 is deprotonated by a distant glutamic acid via the relay action of His-137 and a water molecule. The cysteine then reacts with the peroxide to give a sulfenic acid that subsequently forms a disulfide bond with Cys-130. The crystal structure of AhpD identifies a new protein fold relevant to members of this protein family in other organisms. The structural details constitute a potential platform for the design of inhibitors of potential utility as antitubercular agents and suggest that AhpD may have disulfide exchange properties of importance in other areas of M. tuberculosis biology.
- Department of Biochemistry and Molecular Biology, University College, Gower Street, London WC1E 6BT, United Kingdom.
Organizational Affiliation: