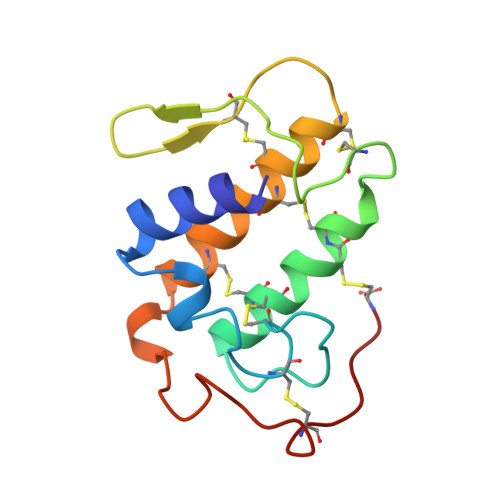
Crystal structure of myotoxin II, a monomeric Lys49-phospholipase A2 homologue isolated from the venom of Cerrophidion (Bothrops) godmani.
Arni, R.K., Fontes, M.R., Barberato, C., Gutierrez, J.M., Diaz, C., Ward, R.J.(1999) Arch Biochem Biophys 366: 177-182
- PubMed: 10356281 Search on PubMed
- DOI: https://doi.org/10.1006/abbi.1999.1210
- Primary Citation Related Structures:
1GOD - PubMed Abstract:
Lys49-Phospholipase A2 (Lys49-PLA2) homologues damage membranes by a Ca2+-independent mechanism which does not involve catalytic activity. With the aim of determining the structural basis for this novel activity, we have solved the crystal structure of myotoxin-II, a Lys49-PLA2 isolated from the venom of Cerrophidion (Bothrops) godmani (godMT-II) at 2.8 A resolution by molecular replacement. The final model has been refined to a final crystallografic residual (Rfactor) of 18.8% (Rfree = 28.2%), with excellent stereochemistry. godMT-II is also monomeric in the crystalline state, and small-angle X-ray scattering results demonstrate that the protein is monomeric in solution under fisicochemical conditions similar to those used in the crystallographic studies.
- Department of Physics, IBILCE/UNESP, São José do Rio Preto-SP, Brazil. arni@df.ibilce.unesp.br
Organizational Affiliation: