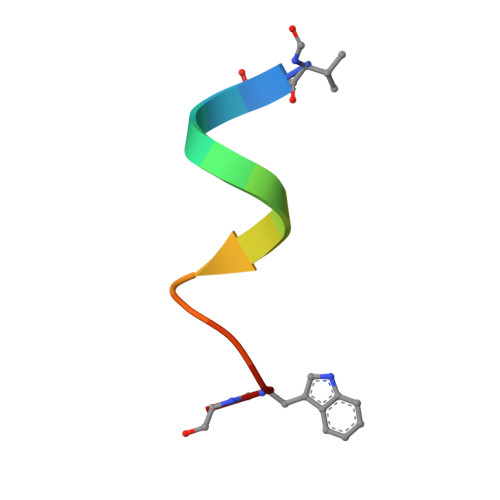
Crystal Structure of the Gramicidin/Potassium Thiocyanate Complex.
Doyle, D.A., Wallace, B.A.(1997) J Mol Biology 266: 963
- PubMed: 9086274 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1996.0837
- Primary Citation Related Structures:
1GMK - PubMed Abstract:
The hydrophobic channel-forming polypeptide gramicidin adopts a left-handed antiparallel double helix conformation with 6.4 residues per turn when in complex with monovalent cation salts in a methanol environment. The crystal structure of the gramicidin/potassium thiocyanate complex (a = 32.06 A, b = 51.80 A, and c = 31.04 A; space group P2(1)2(1)2(1)) has been solved to 2.5 A with an R-factor of 0.193. In the structure, binding sites for the cations are formed by the polypeptide backbone carbonyl groups tilting away from the helix axis toward the ions located in the central lumen. The polypeptide backbone conformations and the side-chain orientations in this potassium complex are significantly different from those in the previously solved gramicidin/caesium chloride crystal complex, due to the requirements for interactions with the smaller sized potassium cation. The locations and numbers of potassium binding sites also differ considerably from the locations and numbers of caesium binding sites in the other structure. Combining information from all the cation binding sites in the two gramicidin/ion complexes produces different views of the three-dimensional structures of a cation as it is transported along a transmembrane pore, and provides an experimental structural basis for modeling the dynamics of peptide-ion binding and ion transport.
- Department of Crystallography, Birkbeck College, University of London, UK.
Organizational Affiliation: