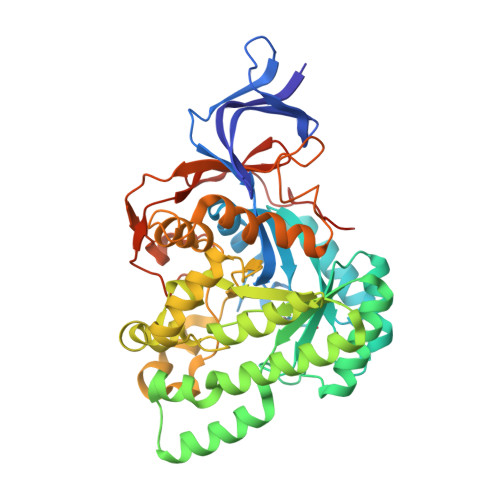
The structure of L-hydantoinase from Arthobacter aurescens leads to an understanding of dihydropyrimidinase substrate and enantio specificity.
Abendroth, J., Niefind, K., May, O., Siemann, M., Syldatk, C., Schomburg, D.(2002) Biochemistry 41: 8589-8597
- PubMed: 12093275 Search on PubMed
- DOI: https://doi.org/10.1021/bi0157722
- Primary Citation Related Structures:
1GKR - PubMed Abstract:
L-Hydantoinase from Arthrobacter aurescens (L-Hyd) is a member of the dihydropyrimidinases which in turn belong to the cyclic amidases. Dihydropyrimidinases catalyze the reversible hydrolytic ring opening of dihydropyrimidines as the second step in the catabolism of pyrimidines. In biotechnology, their hydroloytic activity on five-membered cyclic diamides (hydantoins) is used in the enantio-specific production of amino acids from racemic hydantoins. L-Hyd differs from most of the other dihydropyrimidinases by an L-enantio specificity and by lacking activity on possible natural substrates such as dihydropyrimidines. In this paper, we describe the three-dimensional structure of L-Hyd which was solved by molecular replacement using a homology model and subsequently refined to 2.6 A resolution. Each subunit of the tetrameric L-Hyd consists of an elliptically distorted (alpha/beta)(8)-barrel domain, which hosts the active site, and a beta-sheet domain. In the active site, a binuclear zinc center activates a water molecule for nucleophilic attack on the substrates' amide bond. L-Hyd shows a strong homology both in fold and in metal coordination in the active site to another dihydropyrimidinase from Thermus sp. (D-hydantoinase) and to a slightly lesser degree to ureases, dihydroorotase and phosphotriesterase. Using the homology to ureases, a model for the transition state was modeled in the active site of L-Hyd and D-hydantoinase. This model could provide an explanation for the different substrate and enantio selectivities of both dihydropyrimidinases.
- Institut für Biochemie, Universität zu Köln, Zülpicher Strasse 47, 50674 Köln, Germany.
Organizational Affiliation: