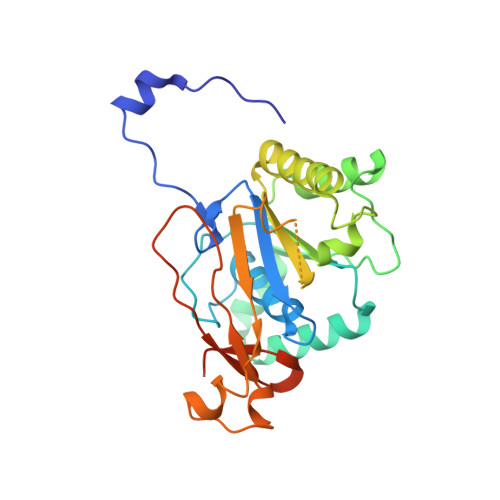
Crystal structure of the hexameric replicative helicase RepA of plasmid RSF1010.
Niedenzu, T., Roleke, D., Bains, G., Scherzinger, E., Saenger, W.(2001) J Mol Biology 306: 479-487
- PubMed: 11178907 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.2000.4398
- Primary Citation Related Structures:
1G8Y - PubMed Abstract:
Unwinding of double-stranded DNA into single-stranded intermediates required for various fundamental life processes is catalyzed by helicases, a family of mono-, di- or hexameric motor proteins fueled by nucleoside triphosphate hydrolysis. The three-dimensional crystal structure of the hexameric helicase RepA encoded by plasmid RSF1010 has been determined by X-ray diffraction at 2.4 A resolution. The hexamer shows an annular structure with 6-fold rotational symmetry and a approximately 17 A wide central hole, suggesting that single-stranded DNA may be threaded during unwinding. Homologs of all five conserved sequence motifs of the DnaB-like helicase family are found in RepA, and the topography of the monomer resembles RecA and the helicase domain of the bacteriophage T7 gp4 protein. In a modeled complex, ATP molecules are located at the subunit interfaces and clearly define adenine-binding and ATPase catalytic sites formed by amino acid residues located on adjacent monomers; most remarkable is the "arginine finger" Arg207 contributing to the active site in the adjacent monomer. This arrangement of active-site residues suggests cooperativity between monomers in ATP hydrolysis and helicase activity of RepA. The mechanism of DNA unwinding remains elusive, as RepA is 6-fold symmetric, contrasting the recently published asymmetric structure of the bacteriophage T7 gp4 helicase domain.
- Institut für Kristallographie, Freie Universität Berlin, Takustr. 6, Berlin, D-14195, Germany.
Organizational Affiliation: