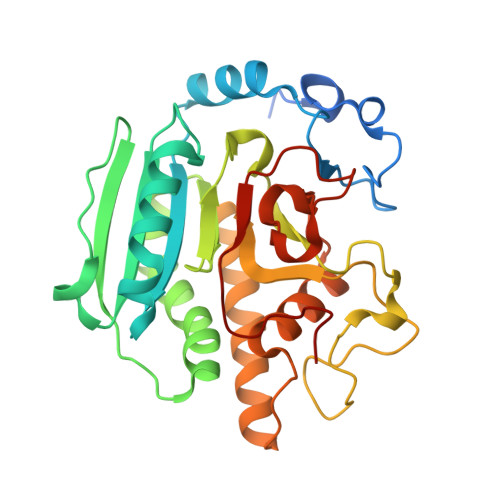
Bovine alpha1,3-galactosyltransferase catalytic domain structure and its relationship with ABO histo-blood group and glycosphingolipid glycosyltransferases.
Gastinel, L.N., Bignon, C., Misra, A.K., Hindsgaul, O., Shaper, J.H., Joziasse, D.H.(2001) EMBO J 20: 638-649
- PubMed: 11179209 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/emboj/20.4.638
- Primary Citation Related Structures:
1FG5, 1G8O, 1G93 - PubMed Abstract:
alpha1,3-galactosyltransferase (alpha3GalT, EC 2.4.1.151) is a Golgi-resident, type II transmembrane protein that transfers galactose from UDP-alpha-galactose to the terminal N:-acetyllactosamine unit of glycoconjugate glycans, producing the Galalpha1,3Galbeta1,4GlcNAc oligosaccharide structure present in most mammalian glycoproteins. Unlike most other mammals, humans and Old World primates do not possess alpha3GalT activity, which is relevant for the hyperacute rejection observed in pig-to-human xenotransplantation. The crystal structure of the catalytic domain of substrate-free bovine alpha3GalT, solved and refined to 2.3 A resolution, has a globular shape with an alpha/beta fold containing a narrow cleft on one face, and shares a UDP-binding domain (UBD) with the recently solved inverting glycosyltransferases. The substrate-bound complex, solved and refined to 2.5 A, allows the description of residues interacting directly with UDP-galactose. These structural data suggest that the strictly conserved residue E317 is likely to be the catalytic nucleophile involved in galactose transfer with retention of anomeric configuration as accomplished by this enzyme. Moreover, the alpha3GalT structure helps to identify amino acid residues that determine the specificities of the highly homologous ABO histo-blood group and glycosphingolipid glycosyltransferases.
- Architecture et Fonction des Macromolécules Biologiques (AFMB), UMR 6098, CNRS and Universités d'Aix-Marseille I and II, 31 Chemin Joseph Aiguier, 13402 Marseille Cedex 20, France. gastinel@afmb.cnrs-mrs.fr
Organizational Affiliation: