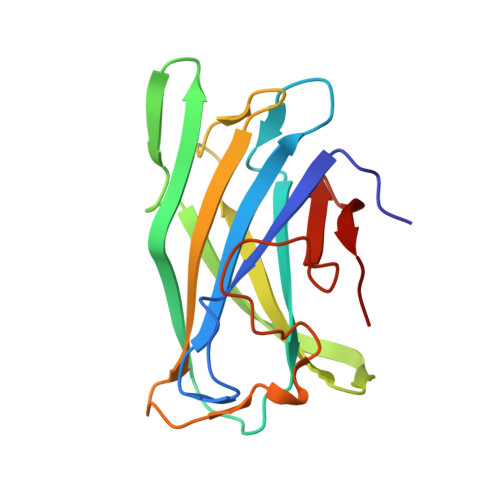
Structure of a family IIIa scaffoldin CBD from the cellulosome of Clostridium cellulolyticum at 2.2 A resolution.
Shimon, L.J., Pages, S., Belaich, A., Belaich, J.P., Bayer, E.A., Lamed, R., Shoham, Y., Frolow, F.(2000) Acta Crystallogr D Biol Crystallogr 56: 1560-1568
- PubMed: 11092922 Search on PubMed
- DOI: https://doi.org/10.1107/s0907444900012889
- Primary Citation Related Structures:
1G43 - PubMed Abstract:
The crystal structure of the family IIIa cellulose-binding domain (CBD) from the cellulosomal scaffoldin subunit (CipC) of Clostridium cellulolyticum has been determined. The structure reveals a nine-stranded jelly-roll topology which exhibits distinctive structural elements consistent with family III CBDs that bind crystalline cellulose. These include a well conserved calcium-binding site, a putative cellulose-binding surface and a conserved shallow groove of unknown function. The CipC CBD structure is very similar to the previously elucidated family IIIa CBD from the CipA scaffoldin of C. thermocellum, with some minor differences. The CipC CBD structure was also compared with other previously described CBD structures from families IIIc and IV derived from the endoglucanases of Thermomonospora fusca and Cellulomonas fimi, respectively. The possible functional consequences of structural similarities and differences in the shallow groove and cellulose-binding faces among various CBD families and subfamilies are discussed.
- Faculty of Chemistry, The Weizmann Institute of Science, Rehovot, Israel. linda.shimon@weizmann.ac.il
Organizational Affiliation: