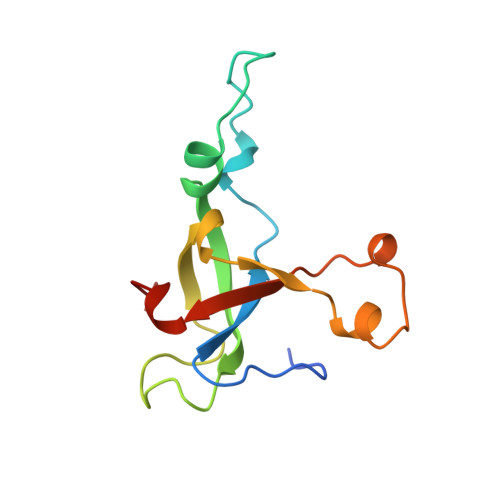
Structural adaptations in the specialized bacteriophage T4 co-chaperonin Gp31 expand the size of the Anfinsen cage.
Hunt, J.F., van der Vies, S.M., Henry, L., Deisenhofer, J.(1997) Cell 90: 361-371
- PubMed: 9244309 Search on PubMed
- DOI: https://doi.org/10.1016/s0092-8674(00)80343-8
- Primary Citation Related Structures:
1G31 - PubMed Abstract:
The Gp31 protein from bacteriophage T4 functionally substitutes for the bacterial co-chaperonin GroES in assisted protein folding reactions both in vitro and in vivo. But Gp31 is required for the folding and/or assembly of the T4 major capsid protein Gp23, and this requirement cannot be satisfied by GroES. The 2.3 A crystal structure of Gp31 shows that its tertiary and quaternary structures are similar to those of GroES despite the existence of only 14% sequence identity between the two proteins. However, Gp31 shows a series of structural adaptations which will increase the size and the hydrophilicity of the "Anfinsen cage," the enclosed cavity within the GroEL/GroES complex that is the location of the chaperonin-assisted protein folding reaction.
- Howard Hughes Medical Institute and Department of Biochemistry, The University of Texas Southwestern Medical Center, Dallas 75235-9050, USA.
Organizational Affiliation: