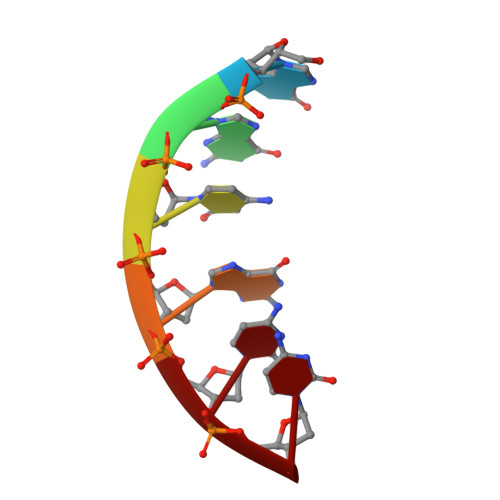
Alternation of DNA and solvent layers in the A form of d(GGCGCC) obtained by ethanol crystallization.
Urpi, L., Navaza, J., Subirana, J.A.(2000) J Biomol Struct Dyn 18: 363-369
- PubMed: 11149513 Search on PubMed
- DOI: https://doi.org/10.1080/07391102.2000.10506673
- Primary Citation Related Structures:
1G00 - PubMed Abstract:
We have determined by X-ray crystallography the structure of the hexamer duplex d(GGCGCC)2 in the A-form using ethanol as a precipitant. The same sequence had previously been crystallized in the B-form, but with 2-methyl-2,4-pentanediol as a precipitant. It appears that ethanol precipitation is a useful method to induce the formation of A-form crystals of DNA. Packing of the molecules in the crystal has unique features: the known interaction of A-DNA duplexes between terminal base-pairs and the minor groove of neighbor molecules is combined with a superstructure consisting in an alternation of DNA layers and solvent layers (water/ions). This organization in layers has been observed before, also with hexamers in the A conformation which crystallize in the same space group (C2221). The solvent layer has a precise thickness, although very few ordered water molecules can be detected. Another feature of this crystal is its large unit cell, which gives rise to an asymmetric unit with three hexamer duplexes. One of the three duplexes is quite different from the other two in several aspects: the number of base pairs per turn, the twist pattern, the mean value of the twist angle and the fact that one terminal base-pair is not stacked as part of the duplex and appears to be disordered. So the variability in conformation of this sequence is remarkable.
- Department d'Enginyeria Química, ETSEIB, Barcelona, Spain.
Organizational Affiliation: