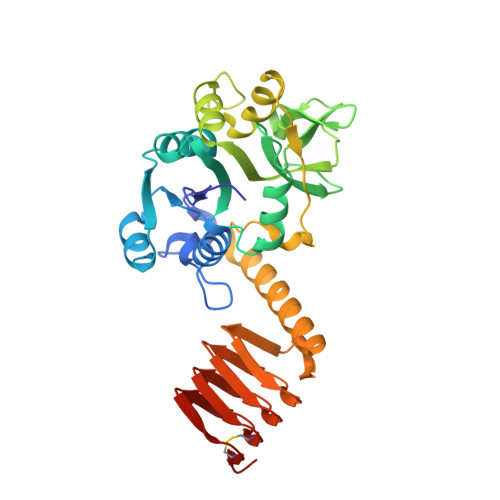
Crystal structure of the bifunctional N-acetylglucosamine 1-phosphate uridyltransferase from Escherichia coli: a paradigm for the related pyrophosphorylase superfamily.
Brown, K., Pompeo, F., Dixon, S., Mengin-Lecreulx, D., Cambillau, C., Bourne, Y.(1999) EMBO J 18: 4096-4107
- PubMed: 10428949 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/emboj/18.15.4096
- Primary Citation Related Structures:
1FWY, 1FXJ - PubMed Abstract:
N-acetylglucosamine 1-phosphate uridyltransferase (GlmU) is a cytoplasmic bifunctional enzyme involved in the biosynthesis of the nucleotide-activated UDP-GlcNAc, which is an essential precursor for the biosynthetic pathways of peptidoglycan and other components in bacteria. The crystal structure of a truncated form of GlmU has been solved at 2.25 A resolution using the multiwavelength anomalous dispersion technique and its function tested with mutagenesis studies. The molecule is composed of two distinct domains connected by a long alpha-helical arm: (i) an N-terminal domain which resembles the dinucleotide-binding Rossmann fold; and (ii) a C-terminal domain which adopts a left-handed parallel beta-helix structure (LbetaH) as found in homologous bacterial acetyltransferases. Three GlmU molecules assemble into a trimeric arrangement with tightly packed parallel LbetaH domains, the long alpha-helical linkers being seated on top of the arrangement and the N-terminal domains projected away from the 3-fold axis. In addition, the 2.3 A resolution structure of the GlmU-UDP-GlcNAc complex reveals the structural bases required for the uridyltransferase activity. These structures exemplify a three-dimensional template for the development of new antibacterial agents and for studying other members of the large family of XDP-sugar bacterial pyrophosphorylases.
- AFMB-CNRS, 31 chemin Joseph Aiguier, 13402 Marseille Cedex 20.
Organizational Affiliation: