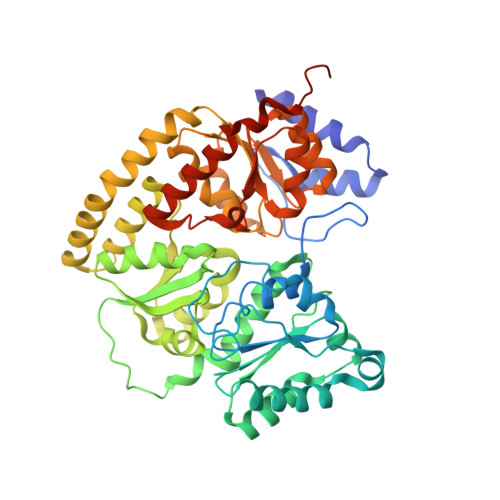
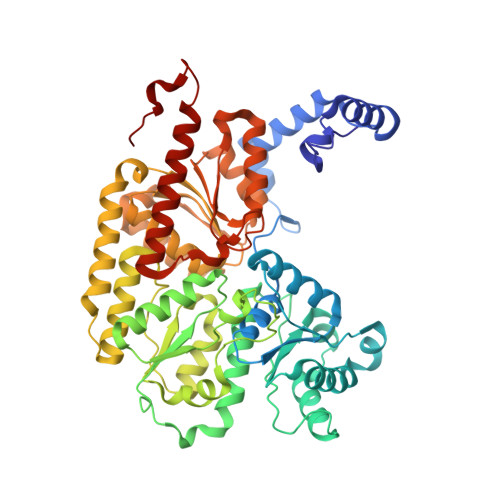
Mechanistic features and structure of the nitrogenase alpha-Gln195 MoFe protein
Sorlie, M., Christiansen, J., Lemon, B.J., Peters, J.W., Dean, D.R., Hales, B.J.(2001) Biochemistry 40: 1540-1549
- PubMed: 11327812 Search on PubMed
- DOI: https://doi.org/10.1021/bi0013997
- Primary Citation Related Structures:
1FP4 - PubMed Abstract:
EPR signals observed under CO and C(2)H(2) during nitrogenase turnover were investigated for the alpha-Gln(195) MoFe protein, an altered form for which the alpha-His(195) residue has been substituted by glutamine. Under CO, samples show S = 1/2 hi- and lo-CO EPR signals identical to those recognized for the wild-type protein, whereas the S = 3/2 signals generated under high CO/high flux conditions differ. Previous work has revealed that the EPR spectrum generated under C(2)H(2) exhibits a signal (S(EPR1)) originating from the FeMo-cofactor having two or more bound C(2)H(2) adducts and a second signal (S(EPR2)) arising from a radical species [Sørlie, M., Christiansen, J., Dean, D. R., and Hales, B. J. (1999) J. Am. Chem. Soc. 121, 9457-9458]. Pressure-dependent studies show that the intensity of these signals has a sigmoidal dependency at low pressures and maximized at 0.1 atm C(2)H(2) with a subsequent decrease in steady-state intensity at higher pressures. Analogous signals are not recognized for the wild-type MoFe protein. Analysis of the principal g-factors of S(EPR2) suggests that it either represents an unusual metal cluster or is a carboxylate centered radical possibly originating from homocitrate. Both S(EPR1) and S(EPR2) exhibit similar relaxation properties that are atypical for S = 1/2 signals originating from Fe-S clusters or radicals and indicate a coupled relaxation pathway. The alpha-Gln(195) MoFe protein also exhibits these signals when incubated under turnover conditions in the presence of C(2)H(4). Under these conditions, additional inflections in the g 4-6 region assigned to ground-state transitions of an S = 3/2 spin system are also recognized and assigned to turnover states of the MoFe protein without C(2)H(4) bound. The structure of alpha-Gln(195) was crystallographically determined and found to be virtually identical to that of the wild-type MoFe protein except for replacement of an NuH-S hydrogen bond interaction between FeMo-cofactor and the imidazole side chain of alpha-His(195) by an analogous interaction involving Gln.
- Department of Chemistry, Louisiana State University, Baton Rouge, LA 70803-1804, USA.
Organizational Affiliation: