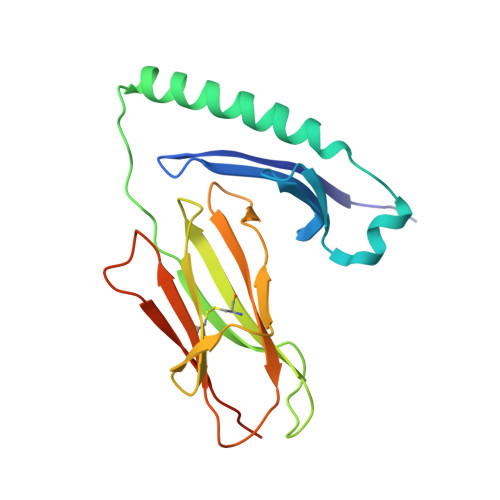
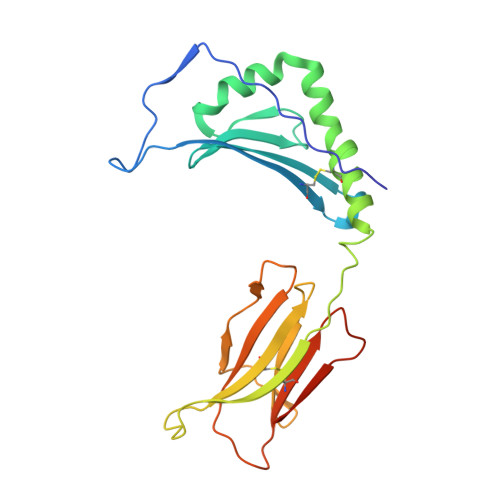
Structural and functional consequences of altering a peptide MHC anchor residue.
Kersh, G.J., Miley, M.J., Nelson, C.A., Grakoui, A., Horvath, S., Donermeyer, D.L., Kappler, J., Allen, P.M., Fremont, D.H.(2001) J Immunol 166: 3345-3354
- PubMed: 11207290 Search on PubMed
- DOI: https://doi.org/10.4049/jimmunol.166.5.3345
- Primary Citation Related Structures:
1FNE, 1FNG - PubMed Abstract:
To better understand TCR discrimination of multiple ligands, we have analyzed the crystal structures of two Hb peptide/I-E(k) complexes that differ by only a single amino acid substitution at the P6 anchor position within the peptide (E73D). Detailed comparison of multiple independently determined structures at 1.9 A resolution reveals that removal of a single buried methylene group can alter a critical portion of the TCR recognition surface. Significant variance was observed in the peptide P5-P8 main chain as well as a rotamer difference at LeuP8, approximately 10 A distal from the substitution. No significant variations were observed in the conformation of the two MHC class II molecules. The ligand alteration results in two peptide/MHC complexes that generate bulk T cell responses that are distinct and essentially nonoverlapping. For the Hb-specific T cell 3.L2, substitution reduces the potency of the ligand 1000-fold. Soluble 3.L2 TCR binds the two peptide/MHC complexes with similar affinity, although with faster kinetics. These results highlight the role of subtle variations in MHC Ag presentation on T cell activation and signaling.
- Department of Pathology and Center for Immunology, Washington University School of Medicine, St. Louis, MO 63110, USA.
Organizational Affiliation: