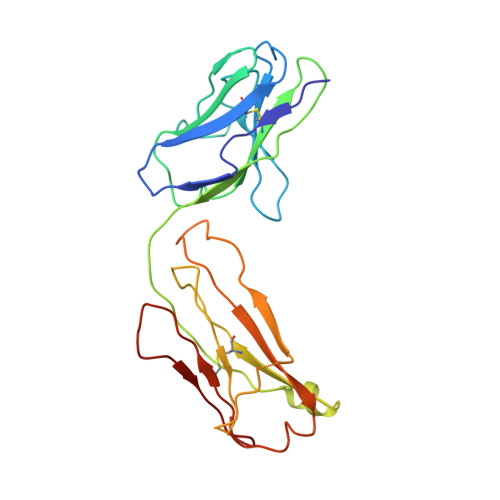
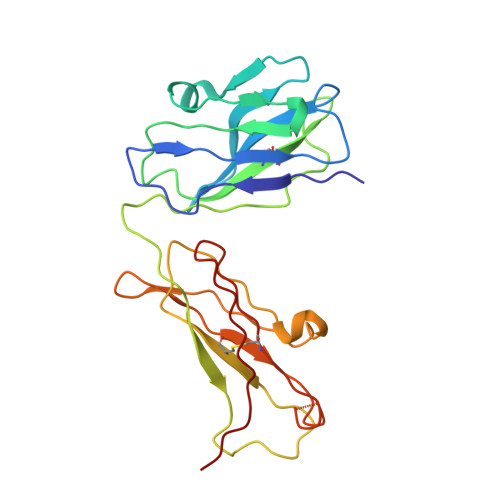
Crystal structure of Fab198, an efficient protector of the acetylcholine receptor against myasthenogenic antibodies.
Poulas, K., Eliopoulos, E., Vatzaki, E., Navaza, J., Kontou, M., Oikonomakos, N., Acharya, K.R., Tzartos, S.J.(2001) Eur J Biochem 268: 3685-3693
- PubMed: 11432734 Search on PubMed
- DOI: https://doi.org/10.1046/j.1432-1327.2001.02274.x
- Primary Citation Related Structures:
1FN4 - PubMed Abstract:
The crystal structure of the Fab fragment of the rat monoclonal antibody 198, with protective activity for the main immunogenic region of the human muscle acetylcholine receptor against the destructive action of myasthenic antibodies, has been determined and refined to 2.8 A resolution by X-ray crystallographic methods. The mouse anti-lysozyme Fab D1.3 was used as a search model in molecular replacement with the AMORE software. The complementarity determining regions (CDR)-L2, CDR-H1 and CDR-H2 belong to canonical groups. Loops CDR-L3, CDR-H2 and CDR-H3, which seem to make a major contribution to binding, were analyzed and residues of potential importance for antigen-binding are examined. The antigen-binding site was found to be a long crescent-shaped crevice. The structure should serve as a model in the rational design of very high affinity humanized mutants of Fab198, appropriate for therapeutic approaches in the model autoimmune disease myasthenia gravis.
- Department of Biochemistry, Hellenic Pasteur Institute, Athens, Greece.
Organizational Affiliation: