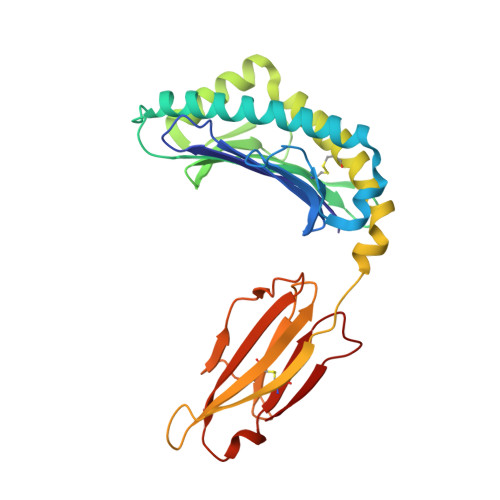
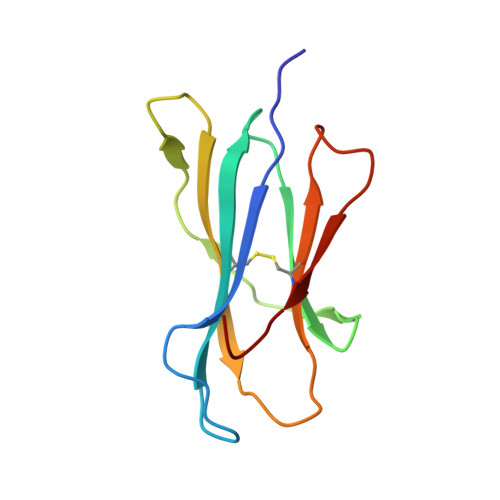
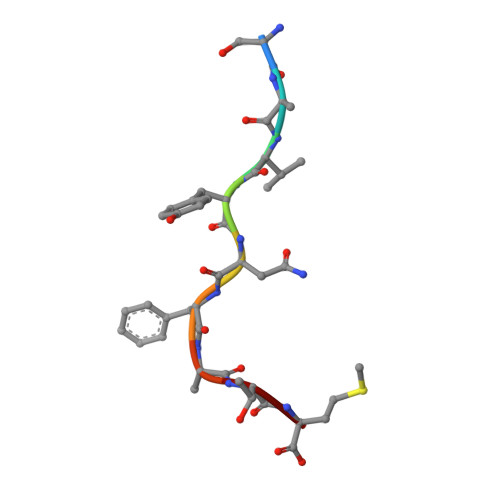
Peptidic termini play a significant role in TCR recognition
Wang, B., Sharma, A., Maile, R., Saad, M., Collins, E.J., Frelinger, J.A.(2002) J Immunol 169: 3137-3145
- PubMed: 12218131 Search on PubMed
- DOI: https://doi.org/10.4049/jimmunol.169.6.3137
- Primary Citation Related Structures:
1FFN, 1FFO, 1FFP - PubMed Abstract:
TCR recognition of class I MHC is dependent on the composition of the antigenic peptide and the MHC. Single amino acid substitutions in either the MHC or the peptide may dramatically alter recognition. While the major interactions between TCR and the peptide/MHC complex appear to be focused on the complementarity-determining region (CDR)3, it is also clear from the cocrystal structure of class I MHC and TCR that the amino and carboxyl ends of the peptide may play a role through interactions with the CDR1. In this work we show that gp33 variants substituted at the peptidic termini at the putative CDR1 contact regions show improved recognition in B6 mice. The rank order of recognition is different using the P14 transgenic T cells, suggesting that one reason for improved recognition is a change in the TCR repertoire that recognizes the peptide. However, the affinity of the TCR by some of the peptide/MHC complex with increased recognition is improved, as shown by increased tetramer binding to P14 T cells. These substitutions at the termini of the peptide-binding cleft cause localized conformational changes as seen by changes in mAb binding and crystallographic structures. The different peptide structures also show different conformations in the center of the peptide, but these are shown to be energetically similar and thus most likely have no significance with respect to TCR recognition. Therefore, small conformational changes, localized to the CDR1 contact regions, may play a significant role in TCR recognition.
- Department of Microbiology and Immunology, University of North Carolina, Chapel Hill, NC 27599, USA.
Organizational Affiliation: