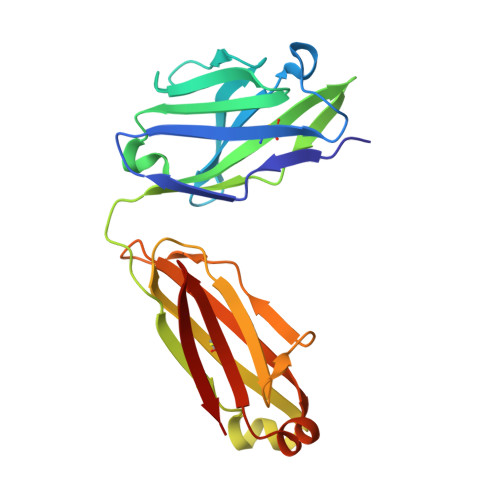
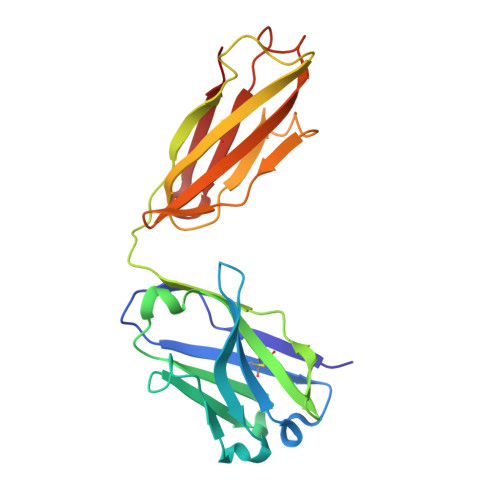
Crystal structure of an anti-carbohydrate antibody directed against Vibrio cholerae O1 in complex with antigen: molecular basis for serotype specificity.
Villeneuve, S., Souchon, H., Riottot, M.M., Mazie, J.C., Lei, P., Glaudemans, C.P., Kovac, P., Fournier, J.M., Alzari, P.M.(2000) Proc Natl Acad Sci U S A 97: 8433-8438
- PubMed: 10880560 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.060022997
- Primary Citation Related Structures:
1F4W, 1F4X, 1F4Y - PubMed Abstract:
The crystal structure of the murine Fab S-20-4 from a protective anti-cholera Ab specific for the lipopolysaccharide Ag of the Ogawa serotype has been determined in its unliganded form and in complex with synthetic fragments of the Ogawa O-specific polysaccharide (O-SP). The upstream terminal O-SP monosaccharide is shown to be the primary antigenic determinant. Additional perosamine residues protrude outwards from the Ab surface and contribute only marginally to the binding affinity and specificity. A complementary water-excluding hydrophobic interface and five Ab-Ag hydrogen bonds are crucial for carbohydrate recognition. The structure reported here explains the serotype specificity of anti-Ogawa Abs and provides a rational basis toward the development of a synthetic carbohydrate-based anti-cholera vaccine.
- Unité de Biochimie Structurale (Centre National de la Recherche Scientifique, Unité de Recherche Associée 2185), Laboratoire d'Ingénierie des Anticorps, Institut Pasteur, Paris, France.
Organizational Affiliation: