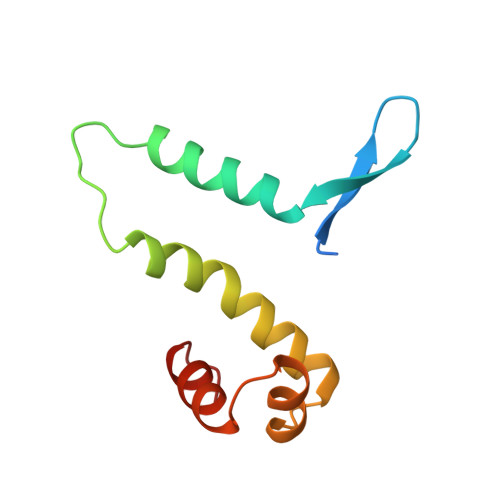
The transactivation region of the fis protein that controls site-specific DNA inversion contains extended mobile beta-hairpin arms.
Safo, M.K., Yang, W.Z., Corselli, L., Cramton, S.E., Yuan, H.S., Johnson, R.C.(1997) EMBO J 16: 6860-6873
- PubMed: 9362499 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/emboj/16.22.6860
- PubMed Abstract:
The Fis protein regulates site-specific DNA inversion catalyzed by a family of DNA invertases when bound to a cis-acting recombinational enhancer. As is often found for transactivation domains, previous crystal structures have failed to resolve the conformation of the N-terminal inversion activation region within the Fis dimer. A new crystal form of a mutant Fis protein now reveals that the activation region contains two beta-hairpin arms that protrude over 20 A from the protein core. Saturation mutagenesis identified the regulatory and structurally important amino acids. The most critical activating residues are located near the tips of the beta-arms. Disulfide cross-linking between the beta-arms demonstrated that they are highly flexible in solution and that efficient inversion activation can occur when the beta-arms are covalently linked together. The emerging picture for this regulatory motif is that contacts with the recombinase at the tip of the mobile beta-arms activate the DNA invertase in the context of an invertasome complex.
- Institute of Molecular Biology, Academia Sinica, Taipei, Taiwan 11529, Republic of China.
Organizational Affiliation: