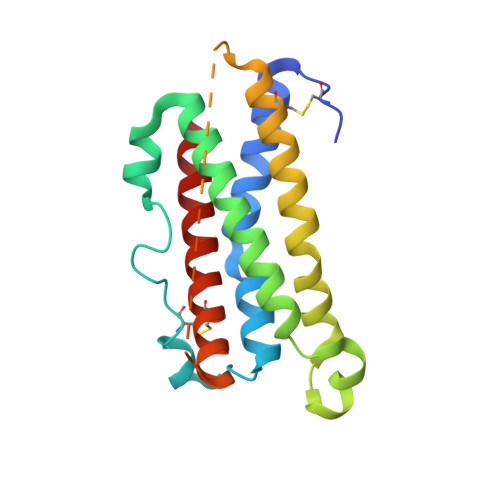
Crystal structure and functional dissection of the cytostatic cytokine oncostatin M.
Deller, M.C., Hudson, K.R., Ikemizu, S., Bravo, J., Jones, E.Y., Heath, J.K.(2000) Structure 8: 863-874
- PubMed: 10997905 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(00)00176-3
- Primary Citation Related Structures:
1EVS - PubMed Abstract:
The cytokine oncostatin M (OSM) inhibits growth of certain tumour-derived cells, induces proliferation in other cell types (e.g. haemangioblasts) and is a mediator of inflammatory responses. Its mechanism of action is via specific binding to gp130 and either the leukaemia inhibitory factor receptor (LIFR) or oncostatin M receptor (OSMR) systems at the cell surface to form an active signalling complex. We report here the crystal structure of human oncostatin M (hOSM) along with mutagenesis data which map the receptor-binding epitopes of the molecule. The structure was determined to a resolution of 2.2 A and conforms to the haematopoietin cytokine up-up-down-down four-helix bundle topology. The site 2 epitope, responsible for gp130 binding, is centred around Gly120 which forms a 'dimple' on the surface of the molecule located on helices A and C. The site 3 motif, responsible for LIFR and OSMR binding, consists of a protruding Phe160/Lys163 pair located at the start of helix D. The data presented allow functional dissection of the receptor-binding interfaces to atomic resolution. Modelling suggests that the gp130 residue Phe169 packs into the site 2 dimple in an analogous fashion to structurally equivalent residues at the growth hormone-growth hormone receptor interface, implying that certain key features may underlie recognition across the whole cytokine/receptor superfamily. Conversely, detailed comparison of the available structures suggests that variations on a common theme dictate the specificity of receptor-ligand interactions within the gp130 family of cytokines.
- Division of Structural Biology, The Wellcome Trust Centre for Human Genetics, University of Oxford, UK.
Organizational Affiliation: