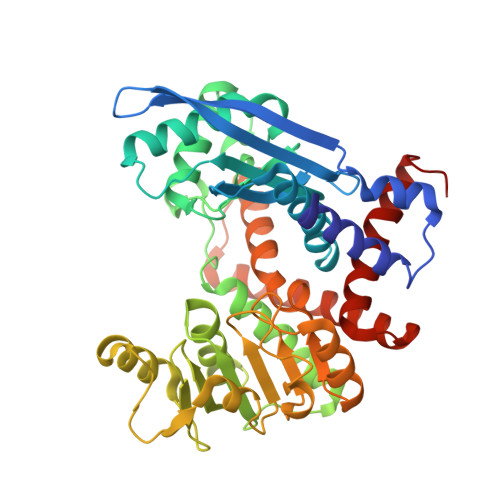
Large-scale domain movements and hydration structure changes in the active-site cleft of unligated glutamate dehydrogenase from Thermococcus profundus studied by cryogenic X-ray crystal structure analysis and small-angle X-ray scattering.
Nakasako, M., Fujisawa, T., Adachi, S., Kudo, T., Higuchi, S.(2001) Biochemistry 40: 3069-3079
- PubMed: 11258921 Search on PubMed
- DOI: https://doi.org/10.1021/bi002482x
- Primary Citation Related Structures:
1EUZ - PubMed Abstract:
Here we describe the large-scale domain movements and hydration structure changes in the active-site cleft of unligated glutamate dehydrogenase. Glutamate dehydrogenase from Thermococcus profundus is composed of six identical subunits of M(r) 46K, each with two distinct domains of roughly equal size separated by a large active-site cleft. The enzyme in the unligated state was crystallized so that one hexamer occupied a crystallographic asymmetric unit, and the crystal structure of the hexamer was solved and refined at a resolution of 2.25 A with a crystallographic R-factor of 0.190. In that structure, the six subunits displayed significant conformational variations with respect to the orientations of the two domains. The variation was most likely explained as a hinge-bending motion caused by small changes in the main chain torsion angle of the residue composing a loop connecting the two domains. Small-angle X-ray scattering profiles both at 293 and 338 K suggested that the apparent molecular size of the hexamer was slightly larger in solution than in the crystalline state. These results led us to the conclusion that (i) the spontaneous domain motion was the property of the enzyme in solution, (ii) the domain motion was trapped in the crystallization process through different modes of crystal contacts, and (iii) the magnitude of the motion in solution was greater than that observed in the crystal structure. The present cryogenic diffraction experiment enabled us to identify 1931 hydration water molecules around the hexamer. The hydration structures around the subunits exhibited significant changes in accord with the degree of the domain movement. In particular, the hydration water molecules in the active-site cleft were rearranged markedly through migrations between specific hydration sites in coupling strongly with the domain movement. We discussed the cooperative dynamics between the domain motion and the hydration structure changes in the active site of the enzyme. The present study provides the first example of a visualized hydration structure varying transiently with the dynamic movements of enzymes and may form a new concept of the dynamics of multidomain enzymes in solution.
- PRESTO, Japan Science and Technology Corporation and The Institute of Molecular and Cellular Biosciences, The University of Tokyo, Yayoi 1-1-1, Bunkyo-ku, Tokyo 113-0032, Japan. tkudo@postman.riken.go.jp
Organizational Affiliation: