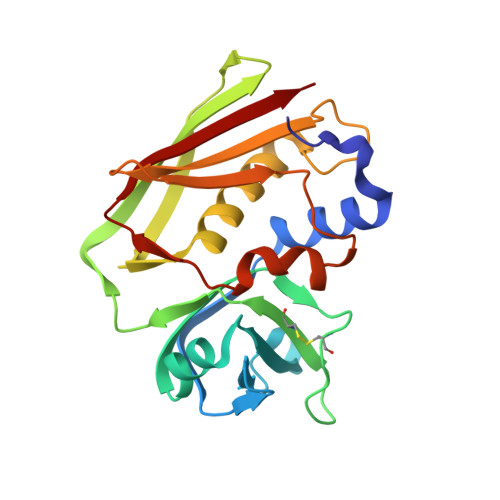
The crystal structure of staphylococcal enterotoxin H: implications for binding properties to MHC class II and TcR molecules.
Hakansson, M., Petersson, K., Nilsson, H., Forsberg, G., Bjork, P., Antonsson, P., Svensson, L.A.(2000) J Mol Biology 302: 527-537
- PubMed: 10986116 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.2000.4093
- Primary Citation Related Structures:
1ENF, 1EWC, 1F77 - PubMed Abstract:
The X-ray structure of the superantigen staphylococcal enterotoxin H (SEH) has been determined at 1.69 A resolution. In this paper we present two structures of zinc-free SEH (apoSEH) and one zinc-loaded form of SEH (ZnSEH). SEH exhibits the conventional superantigen (SAg) fold with two characteristic domains. In ZnSEH one zinc ion per SEH molecule is bound to the C-terminal beta-sheet in the region implicated for major histocompatibility complex class II (MHC class II) binding in SEA, SED and SEE. Surprisingly, the zinc ion has only two ligating amino acid residues His206 and Asp208. The other ligands to the zinc ion are two water molecules. An extensive packing interaction between two symmetry-related molecules in the crystal, 834 A(2)/molecule, forms a cavity that buries the zinc ions of the molecules. This dimer-like interaction is found in two crystal forms. Nevertheless, zinc-dependent dimerisation is not observed in solution, as seen in the case of SED. A unique feature of SEH as compared to other staphylococcal enterotoxins is a large negatively charged surface close to the Zn(2+) site. The interaction of SEH with MHC class II is the strongest known among the staphylococcal enterotoxins. However, SEH seems to lack a SEB-like MHC class II binding site, since the side-chain properties of structurally equivalent amino acid residues in SEH and those in SEB-binding MHC class II differ dramatically. There is also a structural flexibility between the domains of SEH. The domains of two apoSEH structures are related by a 5 degrees rotation leading to at most 3 A difference in C(alpha) positions. Since the T-cell receptor probably interacts with both domains, SEH by this rotation may modulate its binding to different TcR Vbeta-chains.
- Molecular Biophysics, Centre for Chemistry and Chemical Engineering, Lund University, Lund, S-221 00, Sweden.
Organizational Affiliation: