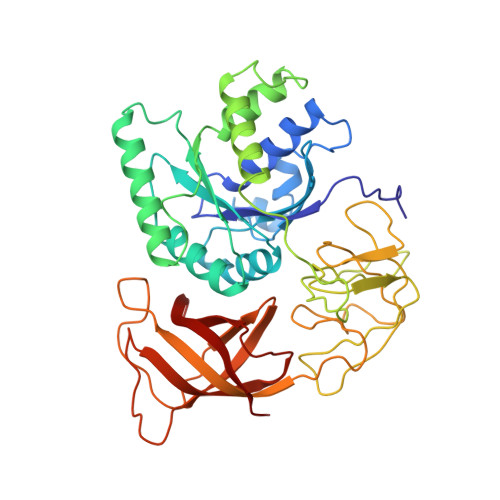
The crystal structure of elongation factor EF-Tu from Thermus aquaticus in the GTP conformation.
Kjeldgaard, M., Nissen, P., Thirup, S., Nyborg, J.(1993) Structure 1: 35-50
- PubMed: 8069622 Search on PubMed
- DOI: https://doi.org/10.1016/0969-2126(93)90007-4
- Primary Citation Related Structures:
1EFT - PubMed Abstract:
Elongation factor Tu (EF-Tu) is a GTP-binding protein that is crucial for protein biosynthesis. In the GTP form of the molecule, EF-Tu binds tightly to aminoacyl-tRNA, forming a ternary complex that interacts with the ribosomal acceptor site. During this interaction, GTP is hydrolyzed, and EF-Tu.GDP is ejected. The crystal structure of EF-Tu from Thermus aquaticus, complexed to the GTP analogue GDPNP, has been determined at 2.5 A resolution and compared to the structure of Escherichia coli EF-Tu.GDP. During the transition from the GDP (inactive) to the GTP (active) form, domain 1, containing the GTP-binding site, undergoes internal conformational changes similar to those observed in ras-p21. In addition, a dramatic rearrangement of domains is observed, corresponding to a rotation of 90.8 degrees of domain 1 relative to domains 2 and 3. Residues that are affected in the binding of aminoacyl-tRNA are found in or near the cleft formed by the domain interface. GTP binding by EF-Tu leads to dramatic conformational changes which expose the tRNA binding site. It appears that tRNA binding to EF-Tu induces a further conformational change, which may affect the GTPase activity.
- Department of Chemistry, Aarhus University, Denmark.
Organizational Affiliation: