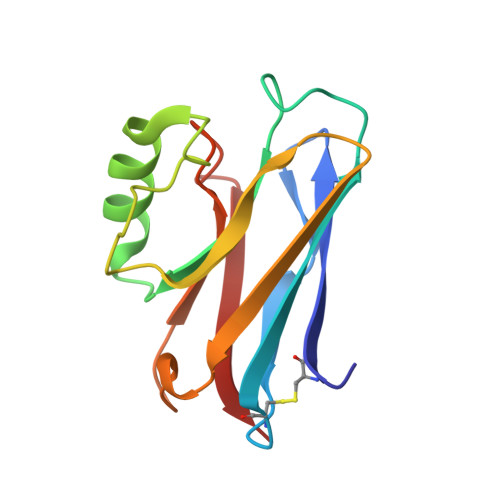
Structures of Oxidised and Reduced Azurin II from Alcaligenes Xylosoxidans at 1.75 Angstoms Resolution
Dodd, F.E., Abraham, Z.H.L., Eady, R.R., Hasnain, S.S.(2000) Acta Crystallogr D Biol Crystallogr 56: 690
- PubMed: 10818345 Search on PubMed
- DOI: https://doi.org/10.1107/s0907444900003309
- Primary Citation Related Structures:
1DYZ, 1DZ0 - PubMed Abstract:
Crystallographic structures of oxidized and reduced forms of azurin II are reported at 1.75 A resolution. Data were collected using one crystal in each case and by translating the crystal after each oscillation range to minimize photoreduction. Very small differences are observed at the Cu site upon reduction and these cannot be determined with confidence at current resolution. A comparison with the three-dimensional EXAFS reveals a good correspondence for all the ligand distances except for Cu-His46, where a larger deviation of approximately 0.12-0.18 A is observed, indicating that this ligand is more tightly restrained in the crystallographic refinement at the current resolution.
- Synchrotron Radiation Department, CCLRC Daresbury Laboratory, Warrington, England.
Organizational Affiliation: