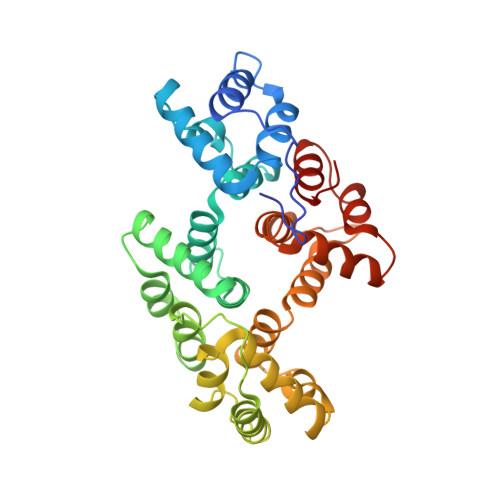
Annexin 24 from Capsicum annuum. X-ray structure and biochemical characterization.
Hofmann, A., Proust, J., Dorowski, A., Schantz, R., Huber, R.(2000) J Biological Chem 275: 8072-8082
- PubMed: 10713128 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.275.11.8072
- Primary Citation Related Structures:
1DK5 - PubMed Abstract:
This work provides the first three-dimensional structure of a member of the plant annexin family and correlates these findings with biochemical properties of this protein. Annexin 24(Ca32) from Capsicum annuum was purified as a native protein from bell pepper and was also prepared by recombinant techniques. To overcome the problem of precipitation of the recombinant wild-type protein in crystallization trials, two mutants were designed. Whereas an N-terminal truncation mutant turned out to be an unstable protein, the N-terminal His-tagged annexin 24(Ca32) was crystallized, and the three-dimensional structure was determined by x-ray diffraction at 2. 8 A resolution. The structure refined to an R-factor of 0.216 adopts the typical annexin fold; the detailed structure, however, is different from non-plant annexins, especially in domains I and III and in the membrane binding loops on the convex side. Within the unit cell there are two molecules per asymmetric unit, which differ in conformation of the IAB-loop. Both conformers show Trp-35 on the surface. The loop-out conformation is stabilized by tight interactions of this tryptophan with residue side chains of a symmetry-related molecule and enforced by a bound sulfate. Characterization of this plant annexin using biophysical methods revealed calcium-dependent binding to phospholipid vesicles with preference for phosphatidylcholine over phosphatidylserine and magnesium-dependent phosphodiesterase activity in vitro as shown with adenosine triphosphate as the substrate. A comparative unfolding study of recombinant annexin 24(Ca32) wild type and of the His-tag fusion protein indicates higher stability of the latter. The effect of this N-terminal modification is also visible from CD spectra. Both proteins were subjected to a FURA-2-based calcium influx assay, which gave high influx rates for the wild-type but greatly reduced influx rates for the fusion protein. We therefore conclude that the N-terminal domain is indeed a major regulatory element modulating different annexin properties by allosteric mechanisms.
- Max-Planck-Institut für Biochemie, Abt. Strukturforschung, 82152 Martinsried, Germany. hofmanna@ncifcrf.gov
Organizational Affiliation: