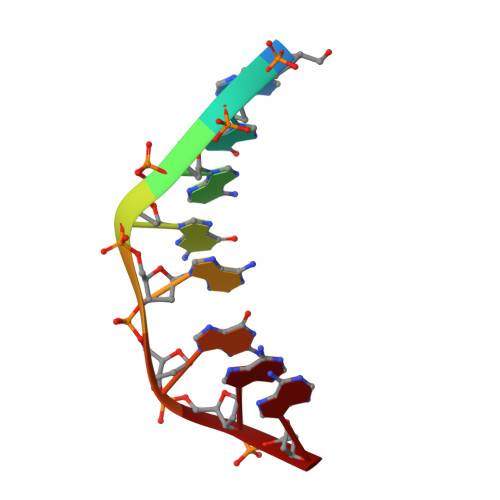
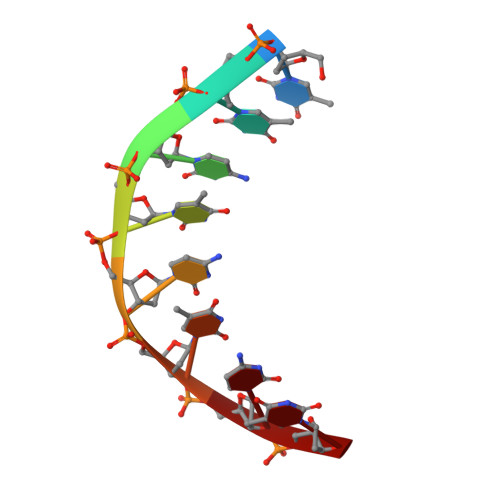
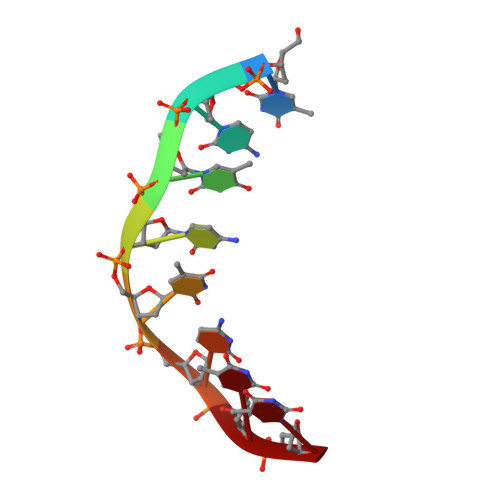
Solution structure of an intramolecular DNA triplex linked by hexakis(ethylene glycol) units: d(AGAGAGAA-(EG)6-TTCTCTCT-(EG)6-TCTCTCTT).
Tarkoy, M., Phipps, A.K., Schultze, P., Feigon, J.(1998) Biochemistry 37: 5810-5819
- PubMed: 9558314 Search on PubMed
- DOI: https://doi.org/10.1021/bi9728102
- Primary Citation Related Structures:
1D3X - PubMed Abstract:
A DNA molecule was designed and synthesized with three octanucleotide stretches linked by two hexakis(ethylene glycol) chains to form an intramolecular triplex in solution. The structural data obtained from a series of NMR NOESY spectra yielded interproton distances, and COSY experiments provided dihedral angle information for analysis of deoxyribose ring pucker. Using distance geometry followed by simulated annealing with restrained molecular dynamics and relaxation matrix refinement, a well-refined ensemble of conformations was calculated. Although some NOE cross-peaks involving protons of the hexakis(ethylene glycol) linker could be identified, most could not be assigned and the conformations of the linkers were not determined. The deoxyribose conformations are predominantly of the S type, except for the protonated cytosine residues in the third strand which show hybrid N and S character. Overall, the duplex part of the molecule resembles a B-DNA double helix with the third strand bound in its major groove by Hoogsteen hydrogen bonds. This structure provides a basis for comparison with triplexes containing noncanonical or nonnatural nucleotides.
- Department of Chemistry and Biochemistry, Molecular Biology Institute, University of California, Los Angeles 90095-1569, USA.
Organizational Affiliation: