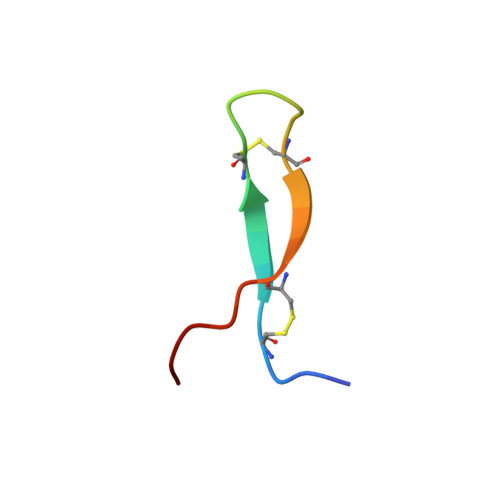
Androctonin, a novel antimicrobial peptide from scorpion Androctonus australis: solution structure and molecular dynamics simulations in the presence of a lipid monolayer.
Mandard, N., Sy, D., Maufrais, C., Bonmatin, J.M., Bulet, P., Hetru, C., Vovelle, F.(1999) J Biomol Struct Dyn 17: 367-380
- PubMed: 10563585 Search on PubMed
- DOI: https://doi.org/10.1080/07391102.1999.10508368
- Primary Citation Related Structures:
1CZ6 - PubMed Abstract:
Androctonin is a highly cationic antimicrobial peptide from scorpion exhibiting a broad spectrum of activities against bacteria and fungi. It contains 25 amino acids including four cysteine residues forming two disulfide bridges. We report here on the determination of its solution structure by conventional two-dimensional (2D) 1H-NMR spectroscopy and molecular modelling using distance geometry and molecular dynamics methods. The structure of androctonin involves a well-defined highly twisted anti-parallel beta-sheet with strands connected by a more variable positively charged turn. A comparison with the structure of tachyplesin I (horseshoe crab) reveals that the amphiphilic character of the protein surface of this homologous peptide is not observed in androctonin. We have undertaken a 200-ps molecular dynamics simulation study on a system including one androctonin molecule and a monolayer of DMPG (1,2-dimyristoylphosphatidylglycerol) lipids. On the basis of this simulation, the first steps of the membrane permeabilization process are discussed.
- Centre de Biophysique Moléculaire, CNRS-UPR 4301, Orléans, France.
Organizational Affiliation: