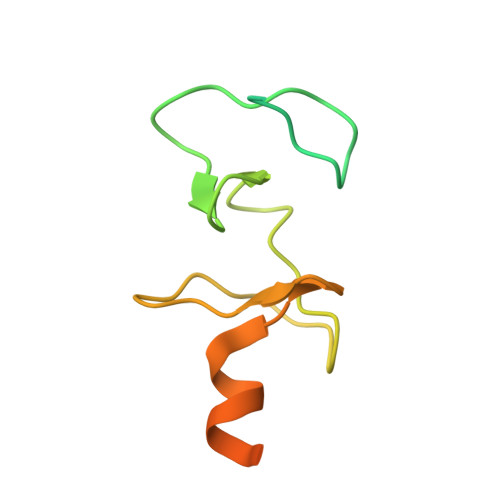
Mutational analysis and NMR spectroscopy of quail cysteine and glycine-rich protein CRP2 reveal an intrinsic segmental flexibility of LIM domains.
Kloiber, K., Weiskirchen, R., Krautler, B., Bister, K., Konrat, R.(1999) J Mol Biology 292: 893-908
- PubMed: 10525413
- DOI: https://doi.org/10.1006/jmbi.1999.3118
- Primary Citation Related Structures:
1CXX - PubMed Abstract:
The LIM domain is a conserved cysteine and histidine-containing structural module of two tandemly arranged zinc fingers. It has been identified in single or multiple copies in a variety of regulatory proteins, either in combination with defined functional domains, like homeodomains, or alone, like in the CRP family of LIM proteins. Structural studies of CRP proteins have allowed a detailed evaluation of interactions in LIM-domains at the molecular level. The packing interactions in the hydrophobic core have been identified as a significant contribution to the LIM domain fold, whereas hydrogen bonding within each single zinc binding site stabilizes zinc finger geometry in a so-called "outer" or "indirect" coordination sphere. Here we report the solution structure of a point-mutant of the carboxyl-terminal LIM domain of quail cysteine and glycine-rich protein CRP2, CRP2(LIM2)R122A, and discuss the structural consequences of the disruption of the hydrogen bond formed between the guanidinium side-chain of Arg122 and the zinc-coordinating cysteine thiolate group in the CCHC rubredoxin-knuckle. The structural analysis revealed that the three-dimensional structure of the CCHC zinc binding site in CRP2(LIM2)R122A is adapted as a consequence of the modified hydrogen bonding pattern. Additionally, as a result of the conformational rearrangement of the zinc binding site, the packing interactions in the hydrophobic core region are altered, leading to a change in the relative orientation of the two zinc fingers with a concomitant change in the solvent accessibilities of hydrophobic residues located at the interface of the two modules. The backbone dynamics of residues located in the folded part of CRP2(LIM2)R122A have been characterized by proton-detected(15)N NMR spectroscopy. Analysis of the R2/R1ratios revealed a rotational correlation time of approximately 6.2 ns and tumbling with an axially symmetric diffusion tensor (D parallel/D perpendicular=1.43). The relaxation data were also analyzed using a reduced spectral density mapping approach. As in wild-type CRP2(LIM2), significant mobility on a picosecond/nanosecond time-scale was detected, and conformational exchange on a microsecond time-scale was identified for residues located in loop regions between secondary structure elements. In summary, the relative orientation of the two zinc binding sites and the accessibility of hydrophobic residues is not only determined by hydrophobic interactions, but can also be modified by the formation and/or breakage of hydrogen bonds. This may be important for the molecular interactions of an adaptor-type LIM domain protein in macromolecular complexes, particularly for the modulation of protein-protein interactions.
- Institute of Organic Chemistry.
Organizational Affiliation: