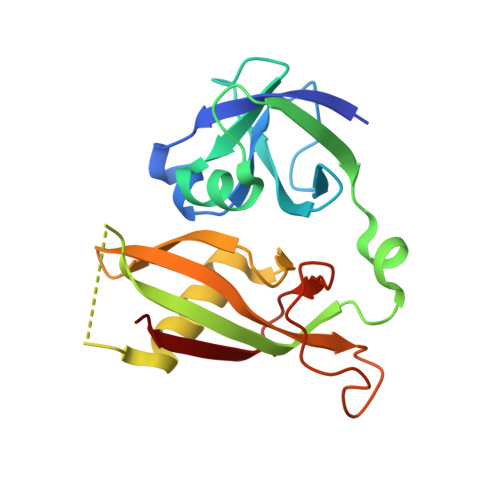
Crystal structure of the Sec18p N-terminal domain.
Babor, S.M., Fass, D.(1999) Proc Natl Acad Sci U S A 96: 14759-14764
- PubMed: 10611286 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.96.26.14759
- Primary Citation Related Structures:
1CR5 - PubMed Abstract:
Yeast Sec18p and its mammalian orthologue N-ethylmaleimide-sensitive fusion protein (NSF) are hexameric ATPases with a central role in vesicle trafficking. Aided by soluble adapter factors (SNAPs), Sec18p/NSF induces ATP-dependent disassembly of a complex of integral membrane proteins from the vesicle and target membranes (SNAP receptors). During the ATP hydrolysis cycle, the Sec18p/NSF homohexamer undergoes a large-scale conformational change involving repositioning of the most N terminal of the three domains of each protomer, a domain that is required for SNAP-mediated interaction with SNAP receptors. Whether an internal conformational change in the N-terminal domains accompanies their reorientation with respect to the rest of the hexamer remains to be addressed. We have determined the structure of the N-terminal domain from Sec18p by x-ray crystallography. The Sec18p N-terminal domain consists of two beta-sheet-rich subdomains connected by a short linker. A conserved basic cleft opposite the linker may constitute a SNAP-binding site. Despite structural variability in the linker region and in an adjacent loop, all three independent molecules in the crystal asymmetric unit have the identical subdomain interface, supporting the notion that this interface is a preferred packing arrangement. However, the linker flexibility allows for the possibility that other subdomain orientations may be sampled.
- Department of Structural Biology, Weizmann Institute of Science, Rehovot 76100, Israel.
Organizational Affiliation: